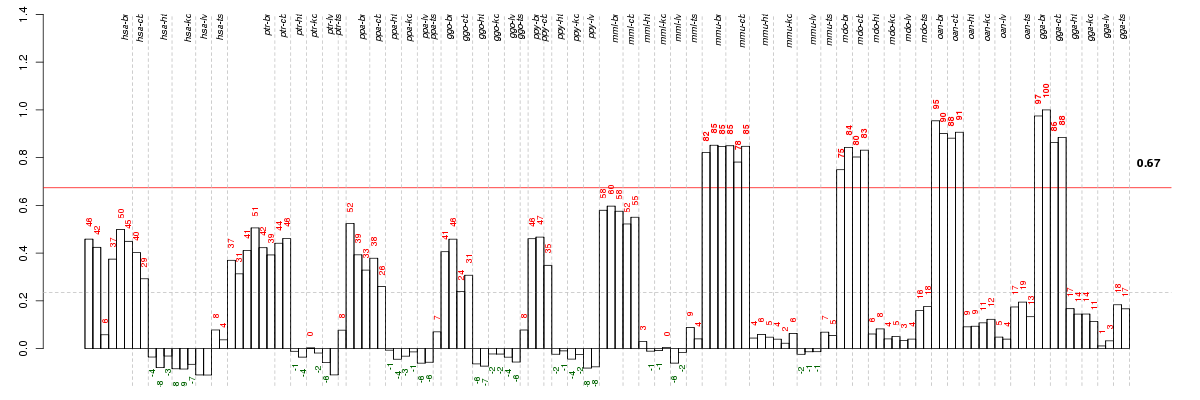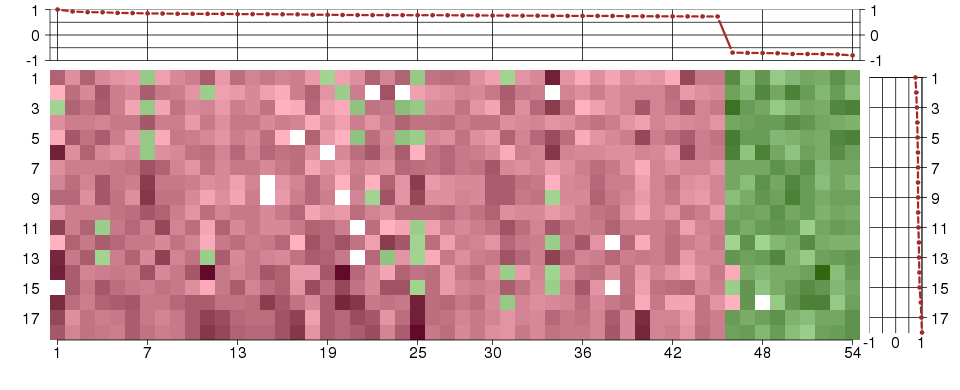

Under-expression is coded with green,
over-expression with red color.


plasma membrane
The membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
membrane
Double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
integral to membrane
Penetrating at least one phospholipid bilayer of a membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer. When used to describe a protein, indicates that all or part of the peptide sequence is embedded in the membrane.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
cell
The basic structural and functional unit of all organisms. Includes the plasma membrane and any external encapsulating structures such as the cell wall and cell envelope.
intrinsic to membrane
Located in a membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
all
NA
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.

molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
catalytic activity
Catalysis of a biochemical reaction at physiological temperatures. In biologically catalyzed reactions, the reactants are known as substrates, and the catalysts are naturally occurring macromolecular substances known as enzymes. Enzymes possess specific binding sites for substrates, and are usually composed wholly or largely of protein, but RNA that has catalytic activity (ribozyme) is often also regarded as enzymatic.
protein kinase activity
Catalysis of the phosphorylation of an amino acid residue in a protein, usually according to the reaction: a protein + ATP = a phosphoprotein + ADP.
protein tyrosine kinase activity
Catalysis of the reaction: ATP + a protein tyrosine = ADP + protein tyrosine phosphate.
transmembrane receptor protein tyrosine kinase activity
Catalysis of the reaction: ATP + a protein-L-tyrosine = ADP + a protein-L-tyrosine phosphate, to initiate a change in cell activity.
signal transducer activity
Mediates the transfer of a signal from the outside to the inside of a cell by means other than the introduction of the signal molecule itself into the cell.
receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity.
transmembrane receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity, and spanning to the membrane of either the cell or an organelle.
ephrin receptor activity
Combining with an ephrin to initiate a change in cell activity.
transporter activity
Enables the directed movement of substances (such as macromolecules, small molecules, ions) into, out of, within or between cells.
amine transmembrane transporter activity
Catalysis of the transfer of amines, including polyamines, from one side of the membrane to the other. Amines are organic compounds that are weakly basic in character and contain an amino (-NH2) or substituted amino group.
amino acid transmembrane transporter activity
Catalysis of the transfer of amino acids from one side of a membrane to the other. Amino acids are organic molecules that contain an amino group and a carboxyl group.
transmembrane transporter activity
Enables the transfer of a substance from one side of a membrane to the other.
organic acid transmembrane transporter activity
Catalysis of the transfer of organic acids, any acidic compound containing carbon in covalent linkage, from one side of the membrane to the other.
L-amino acid transmembrane transporter activity
Catalysis of the transfer of an L-amino acid from one side of a membrane to the other. L-amino acids are the L-enantiomers of amino acids.
L-gamma-aminobutyric acid transmembrane transporter activity
Catalysis of the transfer of L-gamma-aminobutyric acid from one side of a membrane to the other. L-gamma-aminobutyric acid is 4-aminobutyrate (GABA).
kinase activity
Catalysis of the transfer of a phosphate group, usually from ATP, to a substrate molecule.
transferase activity
Catalysis of the transfer of a group, e.g. a methyl group, glycosyl group, acyl group, phosphorus-containing, or other groups, from one compound (generally regarded as the donor) to another compound (generally regarded as the acceptor). Transferase is the systematic name for any enzyme of EC class 2.
transferase activity, transferring phosphorus-containing groups
Catalysis of the transfer of a phosphorus-containing group from one compound (donor) to another (acceptor).
phosphotransferase activity, alcohol group as acceptor
Catalysis of the transfer of a phosphorus-containing group from one compound (donor) to an alcohol group (acceptor).
transmembrane receptor protein kinase activity
NA
active transmembrane transporter activity
Catalysis of the transfer of a specific substance or related group of substances from one side of a membrane to the other, up the solute's concentration gradient. The transporter binds the solute and undergoes a series of conformational changes. Transport works equally well in either direction.
substrate-specific transmembrane transporter activity
Enables the transfer of a specific substance or group of related substances from one side of a membrane to the other.
substrate-specific transporter activity
Enables the directed movement of a specific substance or group of related substances (such as macromolecules, small molecules, ions) into, out of, within or between cells.
carboxylic acid transmembrane transporter activity
Catalysis of the transfer of carboxylic acids from one side of the membrane to the other. Carboxylic acids are organic acids containing one or more carboxyl (COOH) groups or anions (COO-).
molecular transducer activity
The molecular function that accepts an input of one form and creates an output of a different form.
all
NA
substrate-specific transmembrane transporter activity
Enables the transfer of a specific substance or group of related substances from one side of a membrane to the other.
amine transmembrane transporter activity
Catalysis of the transfer of amines, including polyamines, from one side of the membrane to the other. Amines are organic compounds that are weakly basic in character and contain an amino (-NH2) or substituted amino group.
protein kinase activity
Catalysis of the phosphorylation of an amino acid residue in a protein, usually according to the reaction: a protein + ATP = a phosphoprotein + ADP.
transmembrane receptor protein kinase activity
NA
amino acid transmembrane transporter activity
Catalysis of the transfer of amino acids from one side of a membrane to the other. Amino acids are organic molecules that contain an amino group and a carboxyl group.
transmembrane receptor protein tyrosine kinase activity
Catalysis of the reaction: ATP + a protein-L-tyrosine = ADP + a protein-L-tyrosine phosphate, to initiate a change in cell activity.
ABHD3abhydrolase domain containing 3 (ENSG00000158201), score: 0.73 ADAM22ADAM metallopeptidase domain 22 (ENSG00000008277), score: 0.73 ANO3anoctamin 3 (ENSG00000134343), score: 0.77 B3GALT5UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 5 (ENSG00000183778), score: 0.85 C18orf55chromosome 18 open reading frame 55 (ENSG00000075336), score: -0.8 CACNG2calcium channel, voltage-dependent, gamma subunit 2 (ENSG00000166862), score: 0.74 CASTcalpastatin (ENSG00000153113), score: -0.7 CDK17cyclin-dependent kinase 17 (ENSG00000059758), score: 0.82 CDS2CDP-diacylglycerol synthase (phosphatidate cytidylyltransferase) 2 (ENSG00000101290), score: 0.87 CHCHD8coiled-coil-helix-coiled-coil-helix domain containing 8 (ENSG00000181924), score: -0.72 CNR1cannabinoid receptor 1 (brain) (ENSG00000118432), score: 0.75 CNTN5contactin 5 (ENSG00000149972), score: 0.79 COL25A1collagen, type XXV, alpha 1 (ENSG00000188517), score: 0.78 CRTAPcartilage associated protein (ENSG00000170275), score: -0.69 CSMD2CUB and Sushi multiple domains 2 (ENSG00000121904), score: 0.75 CTNND1catenin (cadherin-associated protein), delta 1 (ENSG00000198561), score: -0.75 DGKIdiacylglycerol kinase, iota (ENSG00000157680), score: 0.86 EPHA3EPH receptor A3 (ENSG00000044524), score: 0.74 EPHA5EPH receptor A5 (ENSG00000145242), score: 0.81 EPHA8EPH receptor A8 (ENSG00000070886), score: 0.78 EPHX4epoxide hydrolase 4 (ENSG00000172031), score: 0.78 FADDFas (TNFRSF6)-associated via death domain (ENSG00000168040), score: -0.75 GAD2glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) (ENSG00000136750), score: 0.82 GPR139G protein-coupled receptor 139 (ENSG00000180269), score: 1 GRID2glutamate receptor, ionotropic, delta 2 (ENSG00000152208), score: 0.8 GRM1glutamate receptor, metabotropic 1 (ENSG00000152822), score: 0.73 IL1RAPL1interleukin 1 receptor accessory protein-like 1 (ENSG00000169306), score: 0.82 KCNJ6potassium inwardly-rectifying channel, subfamily J, member 6 (ENSG00000157542), score: 0.9 KIAA0391KIAA0391 (ENSG00000100890), score: -0.75 KIAA2022KIAA2022 (ENSG00000050030), score: 0.78 LGI2leucine-rich repeat LGI family, member 2 (ENSG00000153012), score: 0.92 LOC100291726similar to family with sequence similarity 70, member A (ENSG00000125355), score: 0.79 LRRN1leucine rich repeat neuronal 1 (ENSG00000175928), score: 0.76 LSM10LSM10, U7 small nuclear RNA associated (ENSG00000181817), score: -0.71 MDGA2MAM domain containing glycosylphosphatidylinositol anchor 2 (ENSG00000139915), score: 0.83 MMP16matrix metallopeptidase 16 (membrane-inserted) (ENSG00000156103), score: 0.73 MYO16myosin XVI (ENSG00000041515), score: 0.84 NTSneurotensin (ENSG00000133636), score: 0.83 PPP3CAprotein phosphatase 3, catalytic subunit, alpha isozyme (ENSG00000138814), score: 0.73 PRCPprolylcarboxypeptidase (angiotensinase C) (ENSG00000137509), score: -0.76 RAB3GAP2RAB3 GTPase activating protein subunit 2 (non-catalytic) (ENSG00000118873), score: 0.75 RGS7BPregulator of G-protein signaling 7 binding protein (ENSG00000186479), score: 0.77 RGS8regulator of G-protein signaling 8 (ENSG00000135824), score: 0.89 SEMA3Asema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3A (ENSG00000075213), score: 0.81 SGCZsarcoglycan, zeta (ENSG00000185053), score: 0.75 SLC32A1solute carrier family 32 (GABA vesicular transporter), member 1 (ENSG00000101438), score: 0.83 SLC5A7solute carrier family 5 (choline transporter), member 7 (ENSG00000115665), score: 0.78 SLC6A11solute carrier family 6 (neurotransmitter transporter, GABA), member 11 (ENSG00000132164), score: 0.75 SYNJ1synaptojanin 1 (ENSG00000159082), score: 0.77 THSD7Athrombospondin, type I, domain containing 7A (ENSG00000005108), score: 0.83 TRPC3transient receptor potential cation channel, subfamily C, member 3 (ENSG00000138741), score: 0.76 TRPC5transient receptor potential cation channel, subfamily C, member 5 (ENSG00000072315), score: 0.78 TSPAN2tetraspanin 2 (ENSG00000134198), score: 0.76 WDFY3WD repeat and FYVE domain containing 3 (ENSG00000163625), score: 0.76
| Id | species | tissue | sex | individual |
|---|---|---|---|---|
| mdo_br_m_ca1 | mdo | br | m | _ |
| mmu_cb_m2_ca1 | mmu | cb | m | 2 |
| mdo_cb_m_ca1 | mdo | cb | m | _ |
| mmu_br_m2_ca1 | mmu | br | m | 2 |
| mdo_cb_f_ca1 | mdo | cb | f | _ |
| mdo_br_f_ca1 | mdo | br | f | _ |
| mmu_br_f_ca1 | mmu | br | f | _ |
| mmu_cb_f_ca1 | mmu | cb | f | _ |
| mmu_cb_m1_ca1 | mmu | cb | m | 1 |
| mmu_br_m1_ca1 | mmu | br | m | 1 |
| gga_cb_m_ca1 | gga | cb | m | _ |
| oan_cb_m_ca1 | oan | cb | m | _ |
| gga_cb_f_ca1 | gga | cb | f | _ |
| oan_br_f_ca1 | oan | br | f | _ |
| oan_cb_f_ca1 | oan | cb | f | _ |
| oan_br_m_ca1 | oan | br | m | _ |
| gga_br_m_ca1 | gga | br | m | _ |
| gga_br_f_ca1 | gga | br | f | _ |