Difference between revisions of "ExpressionView"
| Line 4: | Line 4: | ||
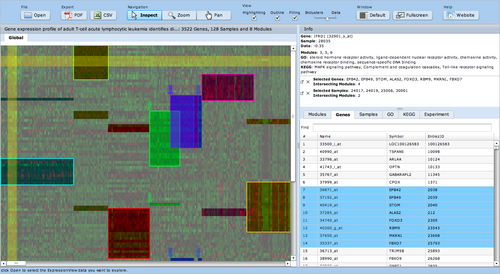
ExpressionView is an R package that provides an interactive environment to explore biclusters identified in gene expression data. A sophisticated ordering algorithm is used to present the biclusters in a visually appealing layout that provides an intuitive summary of the results. From this overview, the user can select individual biclusters and access all the biologically relevant data associated with it. The package is aimed to facilitate the collaboration between bioinformaticians and life scientists who are not familiar with the R language. | ExpressionView is an R package that provides an interactive environment to explore biclusters identified in gene expression data. A sophisticated ordering algorithm is used to present the biclusters in a visually appealing layout that provides an intuitive summary of the results. From this overview, the user can select individual biclusters and access all the biologically relevant data associated with it. The package is aimed to facilitate the collaboration between bioinformaticians and life scientists who are not familiar with the R language. | ||
| − | = Launch = | + | = Launch and download = |
| + | |||
| + | == Launch the ExpressionView Flash applet | ||
* [http://maya:7575/ExpressionView/bin/ExpressionView.html Launch ExpressionView] | * [http://maya:7575/ExpressionView/bin/ExpressionView.html Launch ExpressionView] | ||
* [http://maya:7575/ExpressionView/bin/ExpressionView.html?filename=../data/ALL.small.xml Launch ExpressionView with ALL data (8 modules)] | * [http://maya:7575/ExpressionView/bin/ExpressionView.html?filename=../data/ALL.small.xml Launch ExpressionView with ALL data (8 modules)] | ||
* [http://maya:7575/ExpressionView/bin/ExpressionView.html?filename=../data/ALL.large.xml Launch ExpressionView with ALL data (108 modules)] | * [http://maya:7575/ExpressionView/bin/ExpressionView.html?filename=../data/ALL.large.xml Launch ExpressionView with ALL data (108 modules)] | ||
| + | |||
| + | == Download the R package (includes the Flash applet)== | ||
| + | |||
| + | == Download the standalone viewer (adobe AIR) == | ||
| + | == Download some sample data == | ||
| + | * [[Media:Expressionview.sampledata.all.small.xml|Gene expression profile of adult T-cell acute lymphocytic leukemia (ALL) with 8 modules]] | ||
| + | * [[Media:Expressionview.sampledata.all.large.xml|Gene expression profile of adult T-cell acute lymphocytic leukemia (ALL) with 108 modules]] | ||
| + | |||
| + | |||
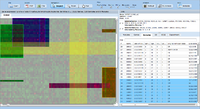
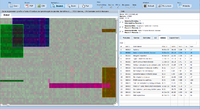
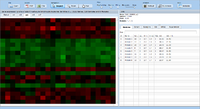
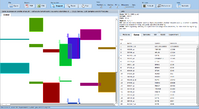
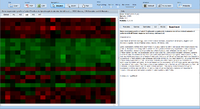
= Screenshots = | = Screenshots = | ||
| Line 21: | Line 32: | ||
image:Expressionview.screenshot.9.png|Experiment description | image:Expressionview.screenshot.9.png|Experiment description | ||
</gallery> | </gallery> | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
= Tutorials = | = Tutorials = | ||
Revision as of 13:46, 16 November 2009
ExpressionView is an R package that provides an interactive environment to explore biclusters identified in gene expression data. A sophisticated ordering algorithm is used to present the biclusters in a visually appealing layout that provides an intuitive summary of the results. From this overview, the user can select individual biclusters and access all the biologically relevant data associated with it. The package is aimed to facilitate the collaboration between bioinformaticians and life scientists who are not familiar with the R language.
Contents
Launch and download
== Launch the ExpressionView Flash applet
- Launch ExpressionView
- Launch ExpressionView with ALL data (8 modules)
- Launch ExpressionView with ALL data (108 modules)
Download the R package (includes the Flash applet)
Download the standalone viewer (adobe AIR)
Download some sample data
- Gene expression profile of adult T-cell acute lymphocytic leukemia (ALL) with 8 modules
- Gene expression profile of adult T-cell acute lymphocytic leukemia (ALL) with 108 modules
Screenshots
Tutorials
There are several tutorials describing how to use ExpressionView: