M2-40 — 13 genes, 6 samples
Enriched GO categories:
None.
Enriched KEGG pathways:
None.
Genes:
AGBL5, C9orf127, DBC1, EGFL6, FGF9, GOLSYN, IGF1, IL27RA, ITIH5, NRCAM, SPON1, STARD5, VAMP1
M2-54 — 12 genes, 5 samples
Enriched GO categories:
None.
Enriched KEGG pathways:
None.
Genes:
AGBL5, C9orf127, DBC1, EGFL6, FGF9, GOLSYN, IL27RA, NME3, NRCAM, SPON1, STARD5, VAMP1
M2-80 — 12 genes, 18 samples
Enriched GO categories:
None.
Enriched KEGG pathways:
None.
Genes:
C15orf5, DBC1, EGFL6, FGF9, FLG, GABRE, GOLSYN, ITIH5, NRCAM, PLGLB2, RALGPS2, SPON1
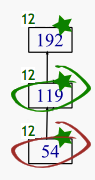
M2-119 — 17 genes, 5 samples
Enriched GO categories:
extracellular region (CC, p=0.020, 7/700)
extracellular region part (CC, p=0.048, 5/387)
Enriched KEGG pathways:
None.
Genes:
AGBL5, C9orf127, CDH8, DBC1, EGFL6, F10, FGF9, GOLSYN, IGF1, IL27RA, ITIH5, NME3, NRCAM, PRELP, SPON1, STARD5, VAMP1
M2-192 — 27 genes, 5 samples
Enriched GO categories:
extracellular region (CC, p=0.0079, 9/700)
extracellular region part (CC, p=0.0403, 6/387)
Enriched KEGG pathways:
None.
Genes:
AGBL5, AKAP6, B3GNT1, BTD, BTN3A2, C9orf127, CDH8, CRYL1, DBC1, EGFL6, F10, FGF9, GOLSYN, GSTT2, HSPB3, IGF1, IL27RA, ITIH5, KLF5, NME3, NRCAM, PLGLB2, PRELP, SPON1, STARD5, VAMP1, VCAM1
M2-284 — 41 genes, 5 samples
Enriched GO categories:
extracellular region (CC, p=0.0057, 12/700)
Enriched KEGG pathways:
None.
Genes:
AGBL5, AKAP6, B3GNT1, BTD, BTN3A2, C9orf127, CDH8, CHST7, CRYL1, CYP26B1, DBC1, EGFL6, F10, FAM13B, FGF9, GABRE, GOLSYN, GSTT2, HSPB3, ID1, IGF1, IL27RA, ITIH5, KLF5, LMOD1, MR1, NME3, NRCAM, PAPPA2, PEX6, …
M2-323 — 32 genes, 8 samples
Enriched GO categories:
extracellular region (CC, p=0.0077, 10/700)
Enriched KEGG pathways:
None.
Genes:
AGBL5, B3GNT1, BTD, BTN3A2, C15orf5, C9orf127, CDH8, DBC1, EGFL6, ENDOD1, F10, FAM13B, FGF9, FLG, GABRE, GOLSYN, HOXA7, HSPB3, IGF1, IL27RA, ITIH5, KLF5, NME3, NRCAM, PAPPA2, PEX6, PLGLB2, PRELP, RALGPS2, SPON1, …
M2-410 — 60 genes, 5 samples
Enriched GO categories:
extracellular region (CC, p=0.011, 14/700)
retinoic acid metabolic process (BP, p=0.044, 2/3)
Enriched KEGG pathways:
None.
Genes:
AGBL5, AKAP6, ALDH1A3, B3GNT1, BTD, BTN3A2, C15orf5, C1orf115, C7orf58, C9orf127, CDH8, CHST7, CRYL1, CYP26B1, DBC1, DDIT3, DNAJC4, EGFL6, ENDOD1, F10, FAM13B, FGF9, FLG, GABRE, GDPD5, GOLSYN, GSTT2, HOXA7, HSPB3, ID1, …
M2-500 — 41 genes, 20 samples
Enriched GO categories:
extracellular region (CC, p=0.014, 11/700)
Enriched KEGG pathways:
None.
Genes:
ABHD14A, ADAMTS6, AGBL5, B3GNT1, BTD, BTN3A2, C15orf5, C1orf115, C9orf127, CHST7, CYP26B1, DBC1, DNAJC4, EGFL6, ENDOD1, F10, FGF9, FLG, GABRE, GDPD5, GOLSYN, HOXA7, IGF1, ITGB8, ITIH5, KLF5, LEPREL2, LMOD1, LPCAT4, NME3, …
M2-607 — 103 genes, 6 samples
Enriched GO categories:
extracellular region (CC, p=0.013, 19/700)
integral to membrane (CC, p=0.032, 37/2022)
intrinsic to membrane (CC, p=0.040, 37/2051)
Enriched KEGG pathways:
None.
Genes:
ABHD14A, ACCN2, AGBL5, AKAP6, ALDH1A3, ARNT2, B3GALT4, B3GNT1, BTD, BTN3A1, BTN3A2, C10orf116, C14orf94, C15orf5, C1orf115, C4orf30, C7orf58, C9orf127, CDH8, CHAC1, CHST7, CRYL1, CYP26B1, DBC1, DDIT3, DNAJC4, DPYSL4, EGFL6, EIF4H, ENDOD1, …
M2-702 — 163 genes, 5 samples
Enriched GO categories:
MHC class I peptide loading complex (CC, p=0.00043, 4/6)
intrinsic to membrane (CC, p=0.00255, 58/2051)
TAP complex (CC, p=0.00273, 3/4)
integral to membrane (CC, p=0.00513, 56/2022)
extracellular region (CC, p=0.00874, 26/700)
Enriched KEGG pathways:
None.
Genes:
ABHD10, ABHD14A, ACCN2, AGBL5, AKAP6, ALDH1A3, ALDH3B1, ARNT2, B3GALT4, B3GNT1, BCL7B, BST1, BTD, BTN3A1, BTN3A2, BTN3A3, C10orf116, C14orf94, C15orf5, C1orf115, C4orf30, C7orf58, C9orf125, C9orf127, CALB2, CAND2, CDH8, CHAC1, CHST7, CLEC3B, …
M2-711 — 155 genes, 6 samples
Enriched GO categories:
extracellular region (CC, p=0.0033, 27/700)
intrinsic to membrane (CC, p=0.0057, 55/2051)
integral to membrane (CC, p=0.0067, 54/2022)
MHC class I peptide loading complex (CC, p=0.0067, 3/6)
membrane part (CC, p=0.0282, 59/2426)
Enriched KEGG pathways:
None.
Genes:
ABHD14A, ACCN2, AGBL5, AKAP6, ALDH1A3, AMPH, ARNT2, B3GALT4, B3GNT1, BAZ1B, BCL7B, BST1, BTD, BTN3A1, BTN3A2, C10orf116, C14orf94, C15orf5, C1orf115, C4orf30, C4orf31, C7orf58, C9orf125, C9orf127, CALB2, CDH8, CHAC1, CHST7, CLEC11A, CLGN, …
M2-741 — 159 genes, 8 samples
Enriched GO categories:
extracellular region (CC, p=0.00086, 30/700)
intrinsic to membrane (CC, p=0.00230, 59/2051)
integral to membrane (CC, p=0.00409, 57/2022)
membrane part (CC, p=0.00642, 64/2426)
MHC class I peptide loading complex (CC, p=0.00711, 3/6)
Enriched KEGG pathways:
None.
Genes:
AGBL5, AKAP6, ALDH1A3, ALDH3B1, ARNT2, B3GNT1, BCL7B, BST1, BTD, BTN3A1, BTN3A2, BTN3A3, C10orf116, C15orf5, C1orf115, C7orf10, C7orf58, C9orf125, C9orf127, CALB2, CDH8, CHST7, CLEC3B, CLGN, COL8A1, CRELD1, CRYL1, CTSC, CTSO, CYP26B1, …
M2-770 — 149 genes, 9 samples
Enriched GO categories:
intrinsic to membrane (CC, p=0.0011, 57/2051)
integral to membrane (CC, p=0.0013, 56/2022)
extracellular region (CC, p=0.0026, 27/700)
membrane part (CC, p=0.0067, 60/2426)
Enriched KEGG pathways:
None.
Genes:
ABCC9, ABHD14A, ACCN2, AGBL5, AKAP6, ALDH1A3, B3GALT4, B3GNT1, BAZ1B, BCL7B, BST1, BTD, BTN3A1, BTN3A2, C10orf116, C14orf94, C15orf5, C1orf115, C4orf31, C7orf58, C9orf127, CDH8, CHST7, CLEC11A, CLEC3B, CLGN, CNTNAP1, CRELD1, CRYBB2, CRYL1, …
M2-806 — 239 genes, 5 samples
Enriched GO categories:
intrinsic to membrane (CC, p=2.1e-05, 89/2051)
integral to membrane (CC, p=3.7e-05, 87/2022)
membrane part (CC, p=8.6e-04, 94/2426)
MHC class I peptide loading complex (CC, p=1.3e-03, 4/6)
extracellular region (CC, p=2.7e-03, 37/700)
Enriched KEGG pathways:
None.
Genes:
ABCC9, ABHD10, ABHD14A, ACCN2, ACVR2A, ADAMTS6, AGBL5, AKAP6, ALDH1A3, ALDH3B1, AMPH, ANKRD28, ARNT2, B3GALT4, B3GNT1, BAZ1B, BCL7B, BST1, BTD, BTN3A1, BTN3A2, BTN3A3, C10orf116, C10orf57, C10orf72, C14orf94, C15orf5, C17orf85, C1orf115, C4orf30, …
M2-809 — 191 genes, 5 samples
Enriched GO categories:
extracellular region (CC, p=0.00019, 36/700)
intrinsic to membrane (CC, p=0.00255, 69/2051)
integral to membrane (CC, p=0.00409, 67/2022)
antigen processing and presentation of peptide antigen (BP, p=0.02167, 5/21)
extracellular region part (CC, p=0.03137, 19/387)
Enriched KEGG pathways:
None.
Genes:
ABCC9, ABHD14A, ADCY9, ADRA2A, AGBL5, AKAP6, ALDH1A3, ALDH1B1, ALDH3B1, ALDH6A1, ALDH9A1, ANKRD28, ARHGEF6, ASPN, ATF4, ATP2A2, B3GNT1, BAG1, BAZ1B, BST1, BTN3A1, BTN3A2, C10orf116, C10orf57, C10orf72, C1orf115, C2orf68, C4orf18, C4orf30, C4orf31, …
M2-827 — 226 genes, 7 samples
Enriched GO categories:
MHC class I peptide loading complex (CC, p=0.0012, 4/6)
intrinsic to membrane (CC, p=0.0013, 78/2051)
integral to membrane (CC, p=0.0014, 77/2022)
TAP complex (CC, p=0.0055, 3/4)
extracellular region (CC, p=0.0060, 34/700)
Enriched KEGG pathways:
None.
Genes:
ABCC9, ABHD10, ABHD14A, ACCN2, ACVR2A, ADAMTS6, AGBL5, AK5, AKAP6, ALDH1A3, AMPH, ARHGEF6, ARNT2, B3GALT4, B3GNT1, BAG1, BAZ1B, BCL7B, BST1, BTD, BTN3A1, BTN3A2, BZW2, C10orf116, C14orf94, C15orf5, C1orf115, C4orf30, C4orf31, C7orf58, …
M2-328 — 12 genes, 14 samples
Enriched GO categories:
None.
Enriched KEGG pathways:
None.
Genes:
CDKN1C, CRTC3, CTDSP2, DHRS3, LOC100132540, LOC339047, LOC399491, PELI2, RERE, ROR2, TBC1D8, ZHX2
M2-379 — 16 genes, 29 samples
Enriched GO categories:
None.
Enriched KEGG pathways:
None.
Genes:
CDKN1C, CRTC3, DMWD, FBXL11, FLJ12529, FNBP4, LHFPL2, LOC100132540, LOC339047, LOC399491, NPIP, PTPN9, RERE, SF1, SLC25A37, TCF20
M2-503 — 73 genes, 28 samples
Enriched GO categories:
extracellular region (CC, p=1.4e-11, 30/700)
extracellular space (CC, p=1.3e-09, 18/246)
extracellular region part (CC, p=2.9e-09, 21/387)
cytokine activity (MF, p=2.7e-08, 11/73)
inflammatory response (BP, p=5.3e-07, 14/153)
Enriched KEGG pathways:
Cytokine-cytokine receptor interaction (0.00075, 10/122)
Genes:
ABCA5, ABCA8, ANKRD1, ANXA3, BMP2, C1orf54, C3, CADPS2, CCL2, CD200, CFH, CFHR1, COL15A1, CREG1, CXCL1, CXCL2, CXCL3, DACT1, DMD, EPHA5, FGF1, FOXN3, HIVEP1, ICAM1, IER3, IFI27, IL1A, IL1B, IL6, IL8, …
M2-852 — 165 genes, 11 samples
Enriched GO categories:
activation of plasma proteins during acute inflammatory response (BP, p=0.048, 4/17)
complement activation (BP, p=0.048, 4/17)
Enriched KEGG pathways:
None.
Genes:
ADAMTS6, AGBL5, AHCTF1, ALDH1A3, ANKRD28, ANKRD36B, ANXA10, APH1B, APOBEC3F, ARL17, ARL2, ARNT2, ASB1, B3GNTL1, B9D1, BAX, BCCIP, BRE, BTD, BTN3A2, C14orf138, C17orf39, C1orf115, C1orf9, C3, C9orf127, CCDC93, CCNT2, CD320, CD55, …
M2-877 — 339 genes, 6 samples
Enriched GO categories:
MHC class I peptide loading complex (CC, p=0.0029, 4/6)
extracellular region (CC, p=0.0029, 47/700)
intrinsic to membrane (CC, p=0.0058, 104/2051)
integral to membrane (CC, p=0.0073, 102/2022)
TAP complex (CC, p=0.0113, 3/4)
Enriched KEGG pathways:
None.
Genes:
ABCC9, ABHD10, ABHD14A, ACCN2, ACVR2A, ACYP1, ADAMTS6, AGBL5, AK5, AKAP6, ALDH1A3, ALDH1B1, ALDH3B1, ALDH6A1, ALDOC, ALG13, AMPH, ANKRD28, APOD, ARHGEF6, ARL6IP5, ARNT2, ARNTL2, ATF4, ATG16L1, B3GALT4, B3GNT1, BAG1, BAZ1B, BCL7B, …
M2-991 — 373 genes, 12 samples
Enriched GO categories:
extracellular region (CC, p=1.8e-12, 73/700)
extracellular region part (CC, p=3.7e-05, 38/387)
immune response (BP, p=2.4e-03, 30/295)
interleukin-1 receptor activity (MF, p=2.7e-03, 4/4)
extracellular space (CC, p=2.9e-03, 24/246)
Enriched KEGG pathways:
None.
Genes:
ABCA1, ABCA5, ABCC3, ABCC9, ABHD10, ABLIM1, ACSL1, ACVR2A, ADAMTS6, ADH1B, ADK, AKAP2, AKR1A1, AKR1C3, ALDH1A3, ALMS1, ANKMY2, ANKRD36B, APOD, ARHGAP26, ARHGEF11, ARPC1B, ATF5, ATG16L1, ATP2B1, BICC1, BMP2K, BRWD1, BTD, C10orf18, …