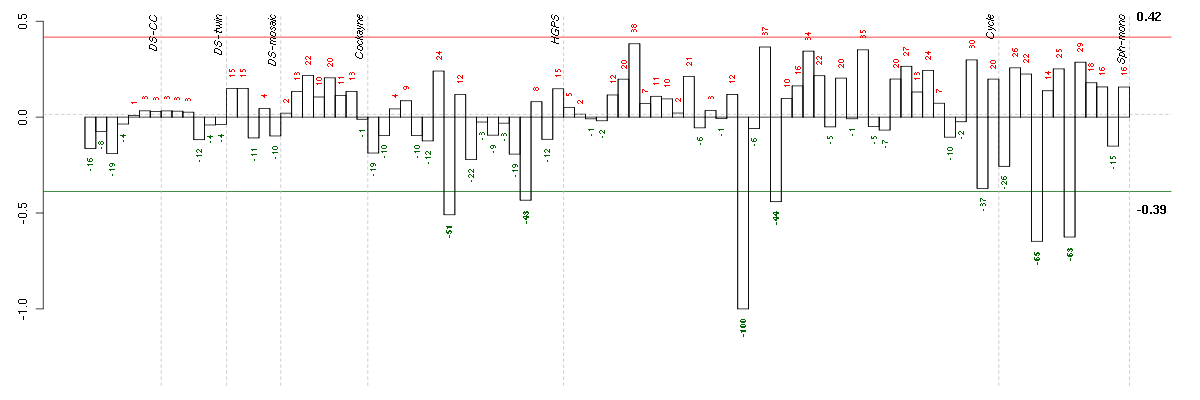

Under-expression is coded with green,
over-expression with red color.



CYTH1cytohesin 1 (202880_s_at), score: -0.72 GIT1G protein-coupled receptor kinase interacting ArfGAP 1 (218030_at), score: -1 IPPKinositol 1,3,4,5,6-pentakisphosphate 2-kinase (219092_s_at), score: -0.85 KCTD2potassium channel tetramerisation domain containing 2 (212564_at), score: -0.7 SDC4syndecan 4 (202071_at), score: -0.71
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485971.cel | 17 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-4219-raw-cel-1311956275.cel | 8 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956398.cel | 12 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-3860-raw-cel-1561690304.cel | 8 | 5 | HGPS | hgu133a | none | GMO8398C |
| E-TABM-263-raw-cel-1515486031.cel | 20 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3860-raw-cel-1561690416.cel | 15 | 5 | HGPS | hgu133a | none | GM0316B |