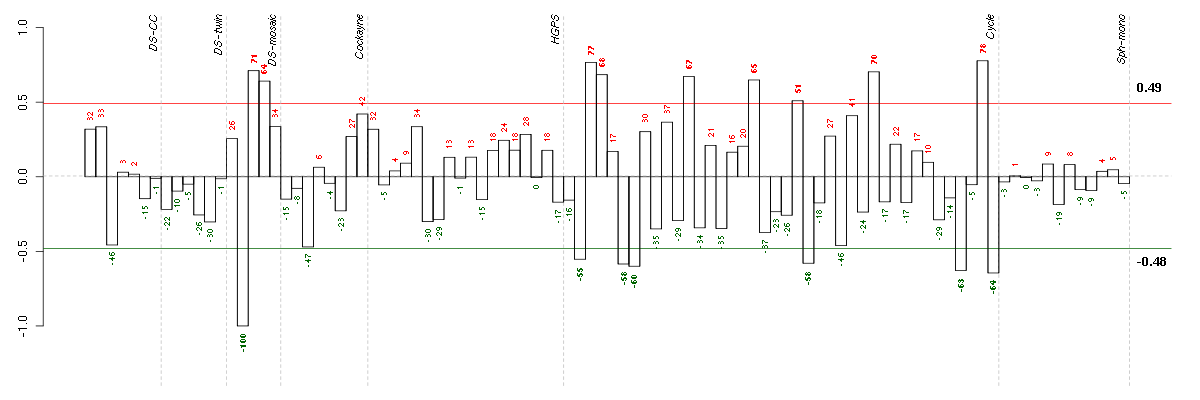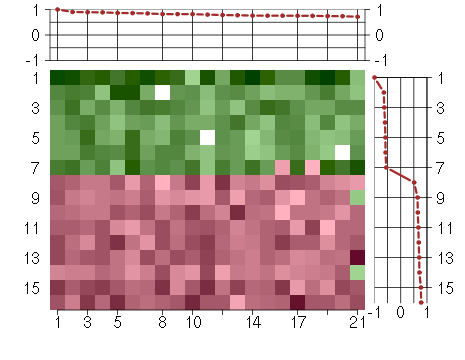

Under-expression is coded with green,
over-expression with red color.



| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 05212 | 4.300e-03 | 0.221 | 4 | 66 | Pancreatic cancer |
| 05220 | 4.991e-02 | 0.2143 | 3 | 64 | Chronic myeloid leukemia |
ADCK2aarF domain containing kinase 2 (221893_s_at), score: 0.75 ARHGDIARho GDP dissociation inhibitor (GDI) alpha (213606_s_at), score: 0.76 BCL2L1BCL2-like 1 (215037_s_at), score: 0.74 BTBD2BTB (POZ) domain containing 2 (207722_s_at), score: 0.82 CNOT3CCR4-NOT transcription complex, subunit 3 (203239_s_at), score: 0.9 DENND3DENN/MADD domain containing 3 (212974_at), score: 0.82 HMGA1high mobility group AT-hook 1 (210457_x_at), score: 0.89 IDUAiduronidase, alpha-L- (205059_s_at), score: 0.78 LRDDleucine-rich repeats and death domain containing (221640_s_at), score: 0.76 NF1neurofibromin 1 (211094_s_at), score: 0.72 NFATC4nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 (205897_at), score: 0.77 PCDHGA11protocadherin gamma subfamily A, 11 (211876_x_at), score: 0.76 PCDHGA3protocadherin gamma subfamily A, 3 (216352_x_at), score: 0.85 PXNpaxillin (211823_s_at), score: 1 RASA3RAS p21 protein activator 3 (206220_s_at), score: 0.75 RELAv-rel reticuloendotheliosis viral oncogene homolog A (avian) (209878_s_at), score: 0.74 SIRT6sirtuin (silent mating type information regulation 2 homolog) 6 (S. cerevisiae) (219613_s_at), score: 0.86 SOLHsmall optic lobes homolog (Drosophila) (204275_at), score: 0.89 TGFBR2transforming growth factor, beta receptor II (70/80kDa) (207334_s_at), score: 0.8 UBTD1ubiquitin domain containing 1 (219172_at), score: 0.87 VEGFBvascular endothelial growth factor B (203683_s_at), score: 0.82
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| 46B.CEL | 2 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 2 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486371.cel | 37 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485771.cel | 7 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485751.cel | 6 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486091.cel | 23 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485671.cel | 2 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 47B.CEL | 4 | 3 | DS-mosaic | hgu133plus2 | Down mosaic | DS-mosaic 4 |
| E-TABM-263-raw-cel-1515485991.cel | 18 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486211.cel | 29 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 46C.CEL | 3 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 3 |
| E-TABM-263-raw-cel-1515485691.cel | 3 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486411.cel | 39 | 6 | Cycle | hgu133a2 | none | Cycle 1 |