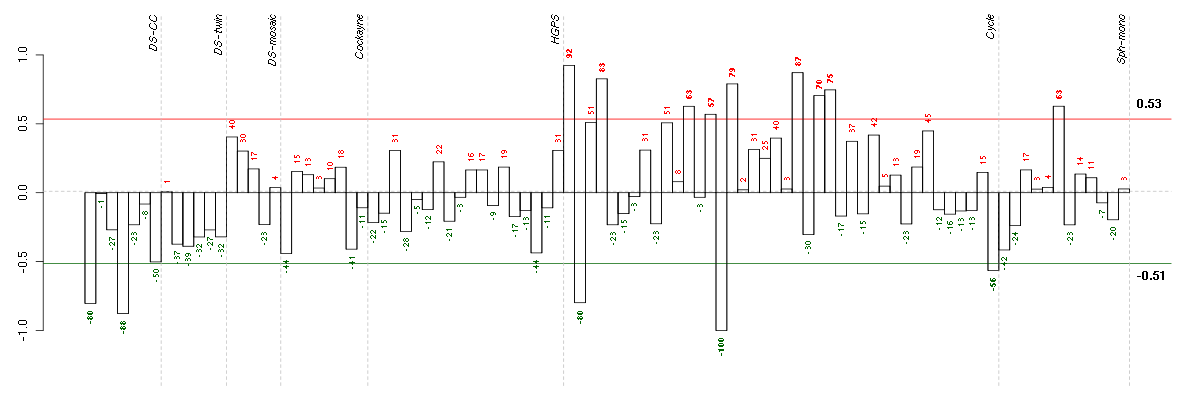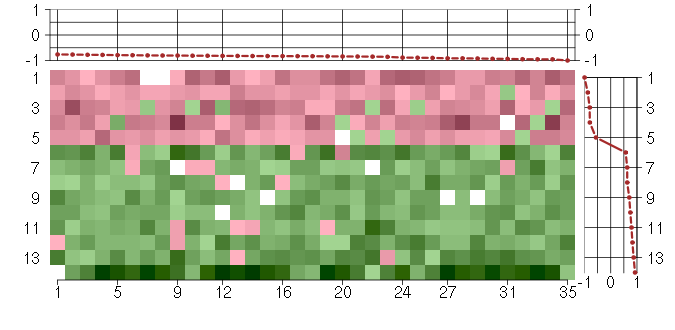

Under-expression is coded with green,
over-expression with red color.



protein binding
Interacting selectively with any protein or protein complex (a complex of two or more proteins that may include other nonprotein molecules).
molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
MAP-kinase scaffold activity
Functions as a physical support for the assembly of a multiprotein mitogen-activated protein kinase (MAPK) complex. MAPK scaffold proteins have binding sites for MAPK pathway kinases as well as for upstream signaling proteins.
structural molecule activity
The action of a molecule that contributes to the structural integrity of a complex or assembly within or outside a cell.
binding
The selective, often stoichiometric, interaction of a molecule with one or more specific sites on another molecule.
receptor signaling complex scaffold activity
Functions to provide a physical support for the assembly of a multiprotein receptor signaling complex.
protein complex scaffold
Functions to provide a physical support for the assembly of a multiprotein complex.
all
This term is the most general term possible
protein complex scaffold
Functions to provide a physical support for the assembly of a multiprotein complex.
ADAMTS7ADAM metallopeptidase with thrombospondin type 1 motif, 7 (220705_s_at), score: -0.92 ADORA1adenosine A1 receptor (216220_s_at), score: -0.9 C21orf2chromosome 21 open reading frame 2 (203996_s_at), score: -0.81 CD53CD53 molecule (203416_at), score: -0.79 COL11A2collagen, type XI, alpha 2 (216993_s_at), score: -0.92 CYP3A4cytochrome P450, family 3, subfamily A, polypeptide 4 (205998_x_at), score: -0.95 FBRSfibrosin (218255_s_at), score: -0.94 FGL1fibrinogen-like 1 (205305_at), score: -0.84 FKBP10FK506 binding protein 10, 65 kDa (219249_s_at), score: -0.95 GPR144G protein-coupled receptor 144 (216289_at), score: -0.85 HAB1B1 for mucin (215778_x_at), score: -1 HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: -0.8 INE1inactivation escape 1 (non-protein coding) (207252_at), score: -0.78 ITGALintegrin, alpha L (antigen CD11A (p180), lymphocyte function-associated antigen 1; alpha polypeptide) (213475_s_at), score: -0.89 LILRB3leukocyte immunoglobulin-like receptor, subfamily B (with TM and ITIM domains), member 3 (211133_x_at), score: -0.89 LOC149478hypothetical protein LOC149478 (215462_at), score: -0.83 LOC80054hypothetical LOC80054 (220465_at), score: -0.91 LRRN2leucine rich repeat neuronal 2 (216164_at), score: -0.77 MAPK8IP1mitogen-activated protein kinase 8 interacting protein 1 (213013_at), score: -0.79 MAPK8IP2mitogen-activated protein kinase 8 interacting protein 2 (205050_s_at), score: -0.81 MOBPmyelin-associated oligodendrocyte basic protein (210193_at), score: -0.82 MUC5ACmucin 5AC, oligomeric mucus/gel-forming (217182_at), score: -0.83 MYL4myosin, light chain 4, alkali; atrial, embryonic (210088_x_at), score: -0.78 NOTCH4Notch homolog 4 (Drosophila) (205247_at), score: -0.84 PGCprogastricsin (pepsinogen C) (205261_at), score: -0.83 PIPOXpipecolic acid oxidase (221605_s_at), score: -0.86 PKLRpyruvate kinase, liver and RBC (222078_at), score: -0.81 PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: -0.84 SEPT5septin 5 (209767_s_at), score: -0.76 SLC26A10solute carrier family 26, member 10 (214951_at), score: -0.8 ST8SIA3ST8 alpha-N-acetyl-neuraminide alpha-2,8-sialyltransferase 3 (208064_s_at), score: -0.95 TBX21T-box 21 (220684_at), score: -0.83 TMSB4Ythymosin beta 4, Y-linked (206769_at), score: -0.93 TRA@T cell receptor alpha locus (216540_at), score: -0.82 TULP2tubby like protein 2 (206733_at), score: -0.82
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485931.cel | 15 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |
| E-TABM-263-raw-cel-1515485671.cel | 2 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485911.cel | 14 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-4219-raw-cel-1311956358.cel | 10 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-TABM-263-raw-cel-1515486111.cel | 24 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486131.cel | 25 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485951.cel | 16 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |