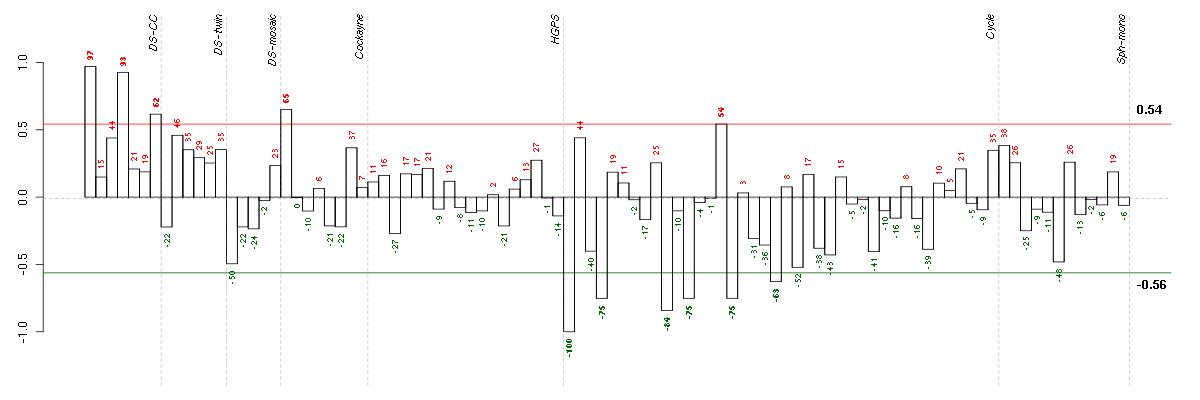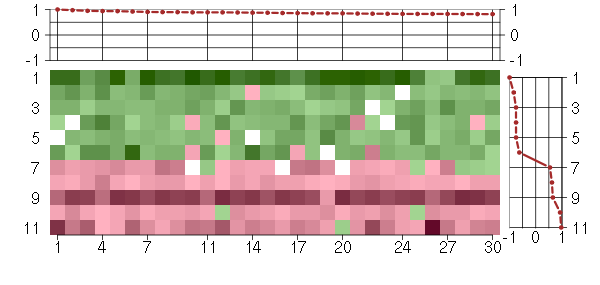

Under-expression is coded with green,
over-expression with red color.



A1CFAPOBEC1 complementation factor (220951_s_at), score: 0.82 BRS3bombesin-like receptor 3 (207369_at), score: 0.83 BTNL3butyrophilin-like 3 (217207_s_at), score: 0.87 C7orf28Achromosome 7 open reading frame 28A (201974_s_at), score: 0.95 C8orf60chromosome 8 open reading frame 60 (220712_at), score: 0.9 COL11A2collagen, type XI, alpha 2 (216993_s_at), score: 0.88 CYP3A4cytochrome P450, family 3, subfamily A, polypeptide 4 (205998_x_at), score: 0.85 DMBT1deleted in malignant brain tumors 1 (208250_s_at), score: 0.83 DNASE1L3deoxyribonuclease I-like 3 (205554_s_at), score: 0.82 DRD5dopamine receptor D5 (208486_at), score: 0.89 F11coagulation factor XI (206610_s_at), score: 0.94 HAB1B1 for mucin (215778_x_at), score: 0.83 HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: 0.85 LHX3LIM homeobox 3 (221670_s_at), score: 0.82 LOH3CR2Aloss of heterozygosity, 3, chromosomal region 2, gene A (220244_at), score: 0.83 LRRN2leucine rich repeat neuronal 2 (216164_at), score: 0.92 MFNGMFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase (204153_s_at), score: 0.84 MOBPmyelin-associated oligodendrocyte basic protein (210193_at), score: 0.89 MYL4myosin, light chain 4, alkali; atrial, embryonic (210088_x_at), score: 0.9 MYO1Amyosin IA (211916_s_at), score: 0.89 PIPOXpipecolic acid oxidase (221605_s_at), score: 0.86 PRSS7protease, serine, 7 (enterokinase) (217269_s_at), score: 0.84 PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: 0.85 RPGRIP1retinitis pigmentosa GTPase regulator interacting protein 1 (206608_s_at), score: 0.83 S100A14S100 calcium binding protein A14 (218677_at), score: 0.9 SLC26A10solute carrier family 26, member 10 (214951_at), score: 0.86 TBX21T-box 21 (220684_at), score: 0.87 TMSB4Ythymosin beta 4, Y-linked (206769_at), score: 0.93 TRA@T cell receptor alpha locus (216540_at), score: 0.97 VGFVGF nerve growth factor inducible (205586_x_at), score: 1
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485951.cel | 16 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486031.cel | 20 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485931.cel | 15 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |