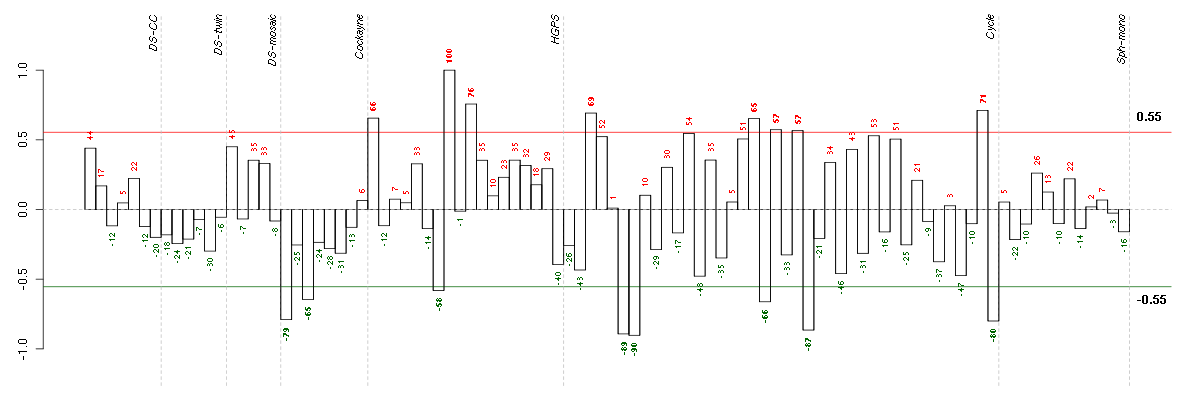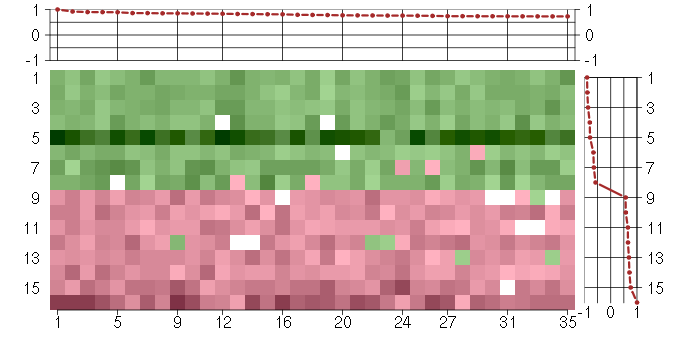

Under-expression is coded with green,
over-expression with red color.



ARSAarylsulfatase A (204443_at), score: 0.76 ATN1atrophin 1 (40489_at), score: 0.82 B3GAT3beta-1,3-glucuronyltransferase 3 (glucuronosyltransferase I) (203452_at), score: 0.81 CDC42BPBCDC42 binding protein kinase beta (DMPK-like) (217849_s_at), score: 0.85 CDR2Lcerebellar degeneration-related protein 2-like (213230_at), score: 0.75 CHST3carbohydrate (chondroitin 6) sulfotransferase 3 (209834_at), score: 0.74 CICcapicua homolog (Drosophila) (212784_at), score: 0.73 DOK4docking protein 4 (209691_s_at), score: 0.74 FAM134Cfamily with sequence similarity 134, member C (212697_at), score: 0.73 FASNfatty acid synthase (212218_s_at), score: 0.73 FBXO17F-box protein 17 (220233_at), score: 0.73 FOXK2forkhead box K2 (203064_s_at), score: 0.82 FURINfurin (paired basic amino acid cleaving enzyme) (201945_at), score: 0.9 GOLGA3golgi autoantigen, golgin subfamily a, 3 (202106_at), score: 0.73 GRINAglutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1 (glutamate binding) (212090_at), score: 0.81 IDUAiduronidase, alpha-L- (205059_s_at), score: 0.77 MAP1Smicrotubule-associated protein 1S (218522_s_at), score: 0.92 MAP7D1MAP7 domain containing 1 (217943_s_at), score: 0.84 PARVBparvin, beta (37965_at), score: 0.86 PCDHG@protocadherin gamma cluster (215836_s_at), score: 0.79 PCDHGA1protocadherin gamma subfamily A, 1 (209079_x_at), score: 0.9 PIP5K1Cphosphatidylinositol-4-phosphate 5-kinase, type I, gamma (212518_at), score: 1 PNPLA6patatin-like phospholipase domain containing 6 (203718_at), score: 0.74 POM121POM121 membrane glycoprotein (rat) (212178_s_at), score: 0.76 PRKACAprotein kinase, cAMP-dependent, catalytic, alpha (202801_at), score: 0.78 SCAMP4secretory carrier membrane protein 4 (213244_at), score: 0.86 SMG7Smg-7 homolog, nonsense mediated mRNA decay factor (C. elegans) (201794_s_at), score: 0.73 SOLHsmall optic lobes homolog (Drosophila) (204275_at), score: 0.76 SSBP3single stranded DNA binding protein 3 (217991_x_at), score: 0.83 STRN4striatin, calmodulin binding protein 4 (217903_at), score: 0.85 SUPT6Hsuppressor of Ty 6 homolog (S. cerevisiae) (208831_x_at), score: 0.77 TAPBPTAP binding protein (tapasin) (208829_at), score: 0.89 TXLNAtaxilin alpha (212300_at), score: 0.79 VEGFBvascular endothelial growth factor B (203683_s_at), score: 0.84 VPS37Bvacuolar protein sorting 37 homolog B (S. cerevisiae) (221704_s_at), score: 0.77
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485771.cel | 7 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485751.cel | 6 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486091.cel | 23 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3407-raw-cel-1437949655.cel | 3 | 4 | Cockayne | hgu133a | none | CSB |
| E-GEOD-3860-raw-cel-1561690272.cel | 7 | 5 | HGPS | hgu133a | HGPS | AG11498 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486031.cel | 20 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485991.cel | 18 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3860-raw-cel-1561690199.cel | 1 | 5 | HGPS | hgu133a | none | GM0316B |
| E-TABM-263-raw-cel-1515485691.cel | 3 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486411.cel | 39 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3860-raw-cel-1561690344.cel | 10 | 5 | HGPS | hgu133a | none | GM00038C |
| E-GEOD-3860-raw-cel-1561690304.cel | 8 | 5 | HGPS | hgu133a | none | GMO8398C |