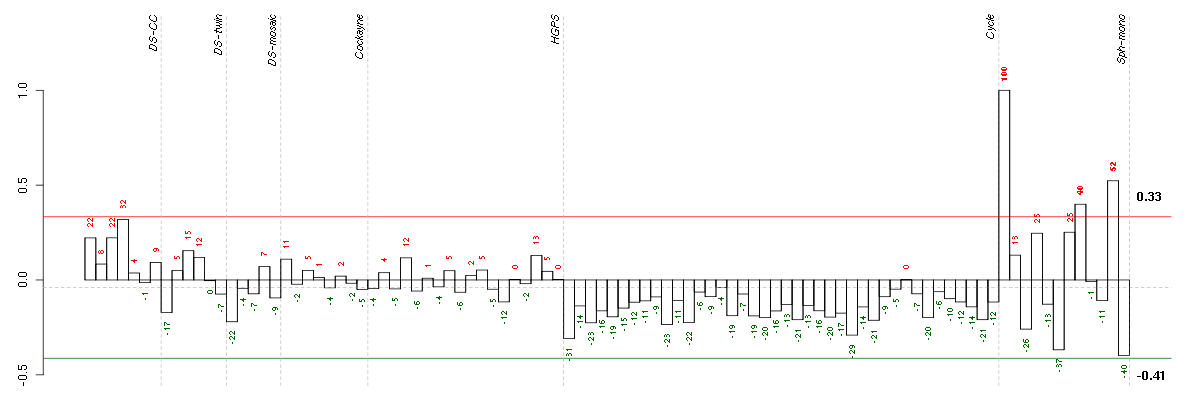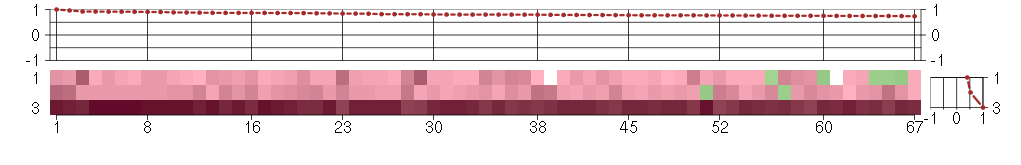

Under-expression is coded with green,
over-expression with red color.



ABCG1ATP-binding cassette, sub-family G (WHITE), member 1 (204567_s_at), score: 0.81 ADORA2Aadenosine A2a receptor (205013_s_at), score: 1 ALMS1Alstrom syndrome 1 (214707_x_at), score: 0.87 ANKRD10ankyrin repeat domain 10 (218093_s_at), score: 0.75 APBB2amyloid beta (A4) precursor protein-binding, family B, member 2 (212972_x_at), score: 0.91 BCL9B-cell CLL/lymphoma 9 (204129_at), score: 0.77 BHLHB9basic helix-loop-helix domain containing, class B, 9 (213709_at), score: 0.87 C4orf10chromosome 4 open reading frame 10 (214123_s_at), score: 0.73 CDC14ACDC14 cell division cycle 14 homolog A (S. cerevisiae) (205288_at), score: 0.79 CDKN1Ccyclin-dependent kinase inhibitor 1C (p57, Kip2) (213183_s_at), score: 0.75 CDR1cerebellar degeneration-related protein 1, 34kDa (207276_at), score: 0.76 CGGBP1CGG triplet repeat binding protein 1 (214050_at), score: 0.89 CHI3L1chitinase 3-like 1 (cartilage glycoprotein-39) (209395_at), score: 0.8 CPEcarboxypeptidase E (201117_s_at), score: 0.8 CXCL2chemokine (C-X-C motif) ligand 2 (209774_x_at), score: 0.81 DAPP1dual adaptor of phosphotyrosine and 3-phosphoinositides (219290_x_at), score: 0.83 DKFZp686O1327hypothetical gene supported by BC043549; BX648102 (216877_at), score: 0.8 DNAH3dynein, axonemal, heavy chain 3 (220725_x_at), score: 0.9 EEF1Deukaryotic translation elongation factor 1 delta (guanine nucleotide exchange protein) (214395_x_at), score: 0.79 FBXW12F-box and WD repeat domain containing 12 (215600_x_at), score: 0.77 FLJ11292hypothetical protein FLJ11292 (220828_s_at), score: 0.91 G3BP1GTPase activating protein (SH3 domain) binding protein 1 (222187_x_at), score: 0.91 GPM6Bglycoprotein M6B (209167_at), score: 0.84 HCG2P7HLA complex group 2 pseudogene 7 (216229_x_at), score: 0.79 HIST2H2BEhistone cluster 2, H2be (202708_s_at), score: 0.77 HNRNPDheterogeneous nuclear ribonucleoprotein D (AU-rich element RNA binding protein 1, 37kDa) (213359_at), score: 0.85 IBD12Inflammatory bowel disease 12 (215373_x_at), score: 0.82 INSRinsulin receptor (213792_s_at), score: 0.86 LOC100128701heterogeneous nuclear ribonucleoprotein A1-like 2 pseudogene (216497_at), score: 0.82 LOC100128836similar to heterogeneous nuclear ribonucleoprotein A1 (217353_at), score: 0.77 LOC643313similar to hypothetical protein LOC284701 (211050_x_at), score: 0.78 LOC644617hypothetical LOC644617 (221235_s_at), score: 0.76 LOC647070hypothetical LOC647070 (215467_x_at), score: 0.76 MCL1myeloid cell leukemia sequence 1 (BCL2-related) (214057_at), score: 0.87 METAP2methionyl aminopeptidase 2 (202015_x_at), score: 0.8 METTL7Amethyltransferase like 7A (211424_x_at), score: 0.8 NACAnascent polypeptide-associated complex alpha subunit (222018_at), score: 0.8 PBX2pre-B-cell leukemia homeobox 2 (202876_s_at), score: 0.8 PELI2pellino homolog 2 (Drosophila) (219132_at), score: 0.87 PHF8PHD finger protein 8 (212916_at), score: 0.75 POGZpogo transposable element with ZNF domain (215281_x_at), score: 0.77 PRINSpsoriasis associated RNA induced by stress (non-protein coding) (216051_x_at), score: 0.78 RAB11FIP1RAB11 family interacting protein 1 (class I) (219681_s_at), score: 0.75 RGS5regulator of G-protein signaling 5 (218353_at), score: 0.92 RPL21P37ribosomal protein L21 pseudogene 37 (216479_at), score: 0.75 RPL23AP32ribosomal protein L23a pseudogene 32 (207283_at), score: 0.78 RPS3AP47ribosomal protein S3a pseudogene 47 (216823_at), score: 0.77 SH3BP2SH3-domain binding protein 2 (217257_at), score: 0.85 SH3GL3SH3-domain GRB2-like 3 (211565_at), score: 0.86 SKP1S-phase kinase-associated protein 1 (200719_at), score: 0.75 SLC30A5solute carrier family 30 (zinc transporter), member 5 (220181_x_at), score: 0.88 SPINLW1serine peptidase inhibitor-like, with Kunitz and WAP domains 1 (eppin) (206318_at), score: 0.86 SPNsialophorin (206057_x_at), score: 0.92 SPTLC3serine palmitoyltransferase, long chain base subunit 3 (220456_at), score: 0.87 SYCP1synaptonemal complex protein 1 (206740_x_at), score: 0.78 TIMM8Atranslocase of inner mitochondrial membrane 8 homolog A (yeast) (210800_at), score: 0.9 TRIM36tripartite motif-containing 36 (219736_at), score: 0.96 TRPV1transient receptor potential cation channel, subfamily V, member 1 (219632_s_at), score: 0.77 UBQLN4ubiquilin 4 (222252_x_at), score: 0.77 UGT2B28UDP glucuronosyltransferase 2 family, polypeptide B28 (211682_x_at), score: 0.75 VAMP2vesicle-associated membrane protein 2 (synaptobrevin 2) (201557_at), score: 0.75 ZBTB39zinc finger and BTB domain containing 39 (205256_at), score: 0.88 ZC3H7Bzinc finger CCCH-type containing 7B (206169_x_at), score: 0.8 ZNF492zinc finger protein 492 (215532_x_at), score: 0.74 ZNF528zinc finger protein 528 (215019_x_at), score: 0.76 ZNF770zinc finger protein 770 (220608_s_at), score: 0.81 ZNF816Azinc finger protein 816A (217541_x_at), score: 0.84
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-GEOD-4219-raw-cel-1311956418.cel | 13 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956634.cel | 19 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956083.cel | 2 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |