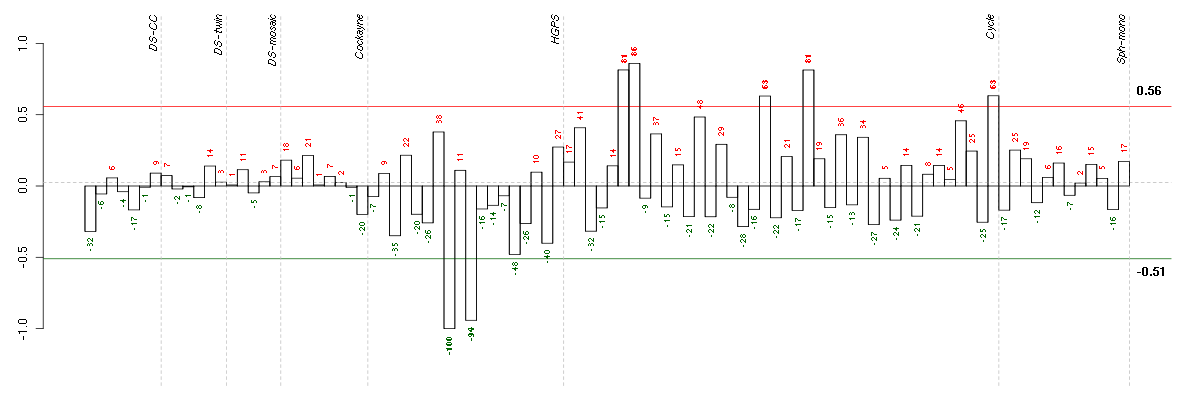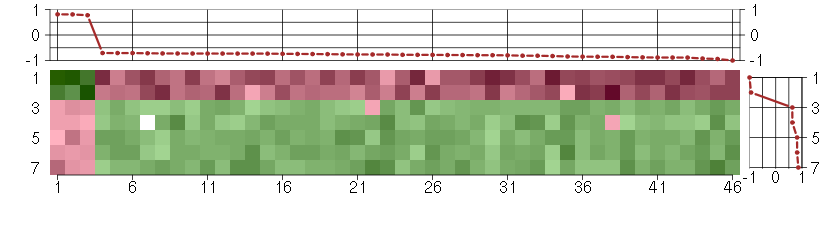

Under-expression is coded with green,
over-expression with red color.



| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 05050 | 4.707e-02 | 0.04688 | 2 | 14 | Dentatorubropallidoluysian atrophy (DRPLA) |
ATN1atrophin 1 (40489_at), score: -0.79 BACE2beta-site APP-cleaving enzyme 2 (217867_x_at), score: -0.78 BAIAP2BAI1-associated protein 2 (205294_at), score: -0.73 BIN1bridging integrator 1 (210201_x_at), score: -0.73 BRD9bromodomain containing 9 (220155_s_at), score: -0.73 CDC42BPBCDC42 binding protein kinase beta (DMPK-like) (217849_s_at), score: -0.71 CDR2Lcerebellar degeneration-related protein 2-like (213230_at), score: -0.76 DCHS1dachsous 1 (Drosophila) (222101_s_at), score: -0.71 DMWDdystrophia myotonica, WD repeat containing (33768_at), score: -0.84 DNM1dynamin 1 (215116_s_at), score: -0.73 EIF2S3eukaryotic translation initiation factor 2, subunit 3 gamma, 52kDa (205321_at), score: 0.77 FAM168Bfamily with sequence similarity 168, member B (212017_at), score: -0.74 FASNfatty acid synthase (212218_s_at), score: -0.76 FLJ12529pre-mRNA cleavage factor I, 59 kDa subunit (217866_at), score: -0.94 FOXK2forkhead box K2 (203064_s_at), score: -0.77 HNRNPH2heterogeneous nuclear ribonucleoprotein H2 (H') (201132_at), score: 0.81 HSPA12Aheat shock 70kDa protein 12A (214434_at), score: -0.76 LOC100132540similar to LOC339047 protein (214870_x_at), score: -0.92 LOC339047hypothetical protein LOC339047 (221501_x_at), score: -1 LOC399491LOC399491 protein (214035_x_at), score: -0.73 LRRN3leucine rich repeat neuronal 3 (209841_s_at), score: 0.81 MAP7D1MAP7 domain containing 1 (217943_s_at), score: -0.75 MAST2microtubule associated serine/threonine kinase 2 (211593_s_at), score: -0.78 MMP14matrix metallopeptidase 14 (membrane-inserted) (202827_s_at), score: -0.72 NBL1neuroblastoma, suppression of tumorigenicity 1 (37005_at), score: -0.88 NFATC4nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 (205897_at), score: -0.73 NKX3-1NK3 homeobox 1 (209706_at), score: -0.79 NPIPnuclear pore complex interacting protein (204538_x_at), score: -0.87 NR1D1nuclear receptor subfamily 1, group D, member 1 (31637_s_at), score: -0.83 OLFM1olfactomedin 1 (213131_at), score: -0.85 OTUD3OTU domain containing 3 (213216_at), score: -0.72 PCDHGA1protocadherin gamma subfamily A, 1 (209079_x_at), score: -0.88 PIP5K1Cphosphatidylinositol-4-phosphate 5-kinase, type I, gamma (212518_at), score: -0.8 POM121POM121 membrane glycoprotein (rat) (212178_s_at), score: -0.81 RNF220ring finger protein 220 (219988_s_at), score: -0.78 SCAMP4secretory carrier membrane protein 4 (213244_at), score: -0.79 SETBP1SET binding protein 1 (205933_at), score: -0.85 SF1splicing factor 1 (208313_s_at), score: -0.75 SIPA1L3signal-induced proliferation-associated 1 like 3 (213600_at), score: -0.74 SLC6A8solute carrier family 6 (neurotransmitter transporter, creatine), member 8 (202219_at), score: -0.71 SMG7Smg-7 homolog, nonsense mediated mRNA decay factor (C. elegans) (201794_s_at), score: -0.73 SOLHsmall optic lobes homolog (Drosophila) (204275_at), score: -0.78 SPON1spondin 1, extracellular matrix protein (209436_at), score: -0.85 STRN4striatin, calmodulin binding protein 4 (217903_at), score: -0.89 TXLNAtaxilin alpha (212300_at), score: -0.81 ZNF536zinc finger protein 536 (206403_at), score: -0.88
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-GEOD-3860-raw-cel-1561690304.cel | 8 | 5 | HGPS | hgu133a | none | GMO8398C |
| E-GEOD-3860-raw-cel-1561690344.cel | 10 | 5 | HGPS | hgu133a | none | GM00038C |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486091.cel | 23 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485751.cel | 6 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485771.cel | 7 | 6 | Cycle | hgu133a2 | none | Cycle 1 |