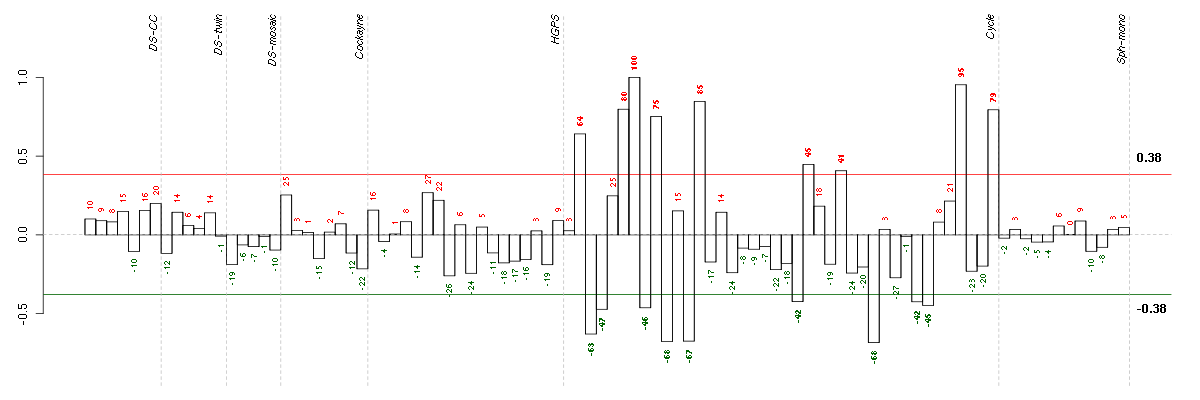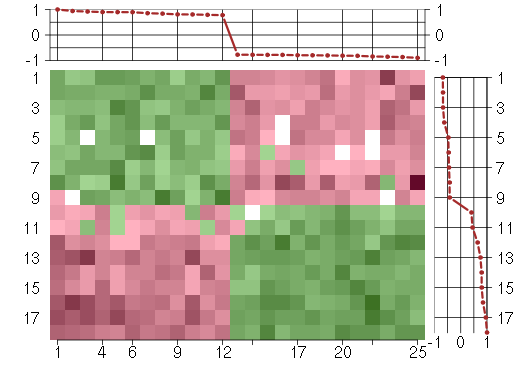

Under-expression is coded with green,
over-expression with red color.



BTBD2BTB (POZ) domain containing 2 (207722_s_at), score: -0.8 CALB2calbindin 2 (205428_s_at), score: 0.89 CNOT3CCR4-NOT transcription complex, subunit 3 (203239_s_at), score: -0.8 CRTC3CREB regulated transcription coactivator 3 (218648_at), score: -0.84 DNAJB9DnaJ (Hsp40) homolog, subfamily B, member 9 (202843_at), score: 0.81 EPHA4EPH receptor A4 (206114_at), score: 0.81 FOXK2forkhead box K2 (203064_s_at), score: -0.81 GFOD1glucose-fructose oxidoreductase domain containing 1 (219821_s_at), score: 0.9 HSD17B6hydroxysteroid (17-beta) dehydrogenase 6 homolog (mouse) (37512_at), score: 0.9 JUPjunction plakoglobin (201015_s_at), score: -0.78 KAL1Kallmann syndrome 1 sequence (205206_at), score: 0.92 KIAA1305KIAA1305 (220911_s_at), score: -0.9 LRP8low density lipoprotein receptor-related protein 8, apolipoprotein e receptor (205282_at), score: 0.85 MYL4myosin, light chain 4, alkali; atrial, embryonic (210088_x_at), score: 0.79 NEFMneurofilament, medium polypeptide (205113_at), score: 1 OLFML2Bolfactomedin-like 2B (213125_at), score: -0.85 PCDH9protocadherin 9 (219737_s_at), score: 0.94 REREarginine-glutamic acid dipeptide (RE) repeats (200940_s_at), score: -0.78 RNF220ring finger protein 220 (219988_s_at), score: -0.79 RPL18AP6ribosomal protein L18a pseudogene 6 (216383_at), score: 0.84 RPS17P5ribosomal protein S17 pseudogene 5 (216348_at), score: 0.78 SCAMP4secretory carrier membrane protein 4 (213244_at), score: -0.87 STIM1stromal interaction molecule 1 (202764_at), score: -0.77 TRIM21tripartite motif-containing 21 (204804_at), score: -0.78 WDR6WD repeat domain 6 (217734_s_at), score: -0.82
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515486211.cel | 29 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485691.cel | 3 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485791.cel | 8 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486311.cel | 34 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486291.cel | 33 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486151.cel | 26 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486091.cel | 23 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485671.cel | 2 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485811.cel | 9 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485751.cel | 6 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485891.cel | 13 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486371.cel | 37 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485771.cel | 7 | 6 | Cycle | hgu133a2 | none | Cycle 1 |