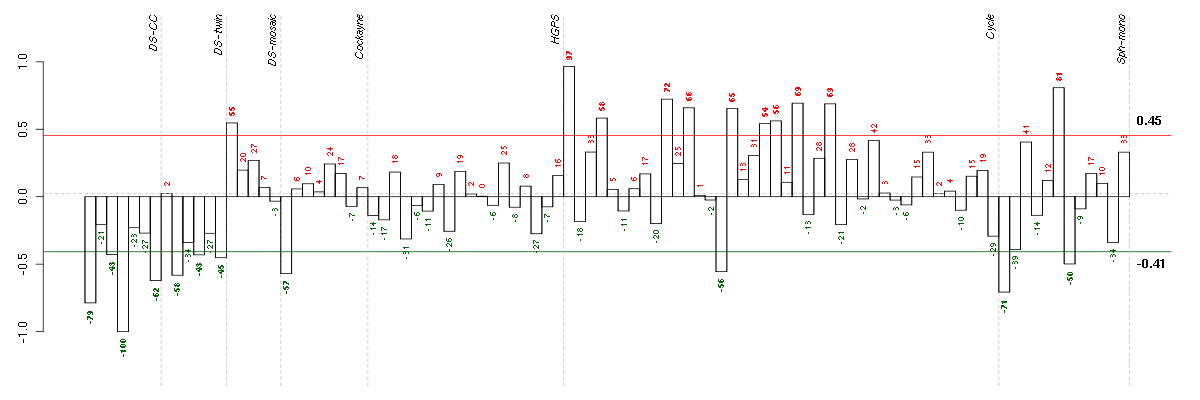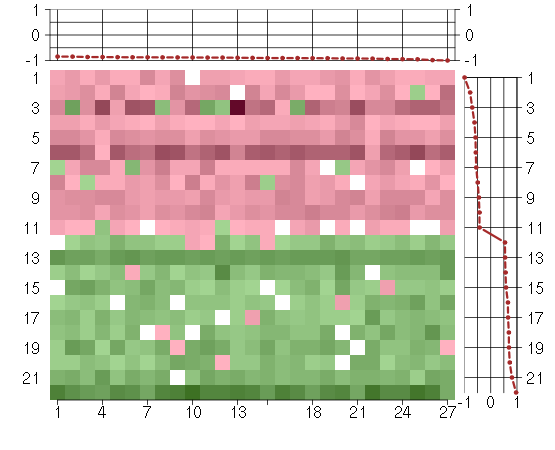

Under-expression is coded with green,
over-expression with red color.



ADORA1adenosine A1 receptor (216220_s_at), score: -0.86 C7orf28Achromosome 7 open reading frame 28A (201974_s_at), score: -1 C8orf60chromosome 8 open reading frame 60 (220712_at), score: -0.88 CHN2chimerin (chimaerin) 2 (207486_x_at), score: -0.91 COL11A2collagen, type XI, alpha 2 (216993_s_at), score: -0.85 CYP3A4cytochrome P450, family 3, subfamily A, polypeptide 4 (205998_x_at), score: -0.93 FAM75A3family with sequence similarity 75, member A3 (215935_at), score: -0.92 FGL1fibrinogen-like 1 (205305_at), score: -0.86 FOLH1folate hydrolase (prostate-specific membrane antigen) 1 (217487_x_at), score: -0.9 HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: -0.95 NCRNA00092non-protein coding RNA 92 (215861_at), score: -0.99 NCRNA00093non-protein coding RNA 93 (210723_x_at), score: -0.95 OR7E24olfactory receptor, family 7, subfamily E, member 24 (215463_at), score: -0.87 PECAM1platelet/endothelial cell adhesion molecule (208982_at), score: -0.92 PRINSpsoriasis associated RNA induced by stress (non-protein coding) (216051_x_at), score: -0.89 PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: -0.89 RP3-377H14.5hypothetical LOC285830 (222279_at), score: -0.88 RP5-886K2.1neuronal thread protein AD7c-NTP (207953_at), score: -0.91 RPGRIP1retinitis pigmentosa GTPase regulator interacting protein 1 (206608_s_at), score: -0.9 S100A14S100 calcium binding protein A14 (218677_at), score: -0.91 SFTPBsurfactant protein B (214354_x_at), score: -0.87 SLC10A1solute carrier family 10 (sodium/bile acid cotransporter family), member 1 (207185_at), score: -0.94 SLC15A1solute carrier family 15 (oligopeptide transporter), member 1 (211349_at), score: -0.85 TBX21T-box 21 (220684_at), score: -0.89 TBX6T-box 6 (207684_at), score: -0.9 TRA@T cell receptor alpha locus (216540_at), score: -0.88
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |
| E-GEOD-4219-raw-cel-1311956083.cel | 2 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| E-TABM-263-raw-cel-1515485931.cel | 15 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-4219-raw-cel-1311956398.cel | 12 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| 6Twin.CEL | 6 | 2 | DS-twin | hgu133plus2 | none | DS-twin 6 |
| 4Twin.CEL | 4 | 2 | DS-twin | hgu133plus2 | none | DS-twin 4 |
| ctrl c 08-03.CEL | 3 | 1 | DS-CC | hgu133a | none | DS-CC 3 |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| E-TABM-263-raw-cel-1515486031.cel | 20 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485951.cel | 16 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486131.cel | 25 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-4219-raw-cel-1311956358.cel | 10 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |