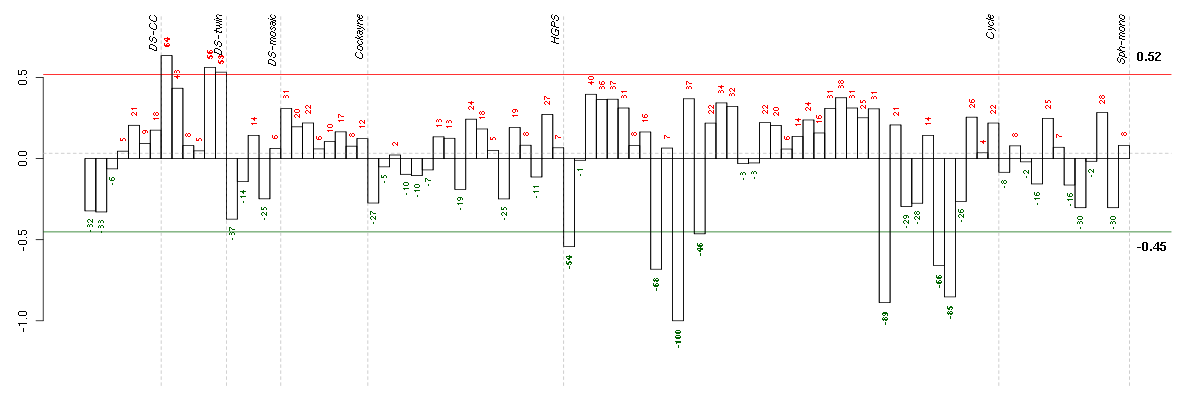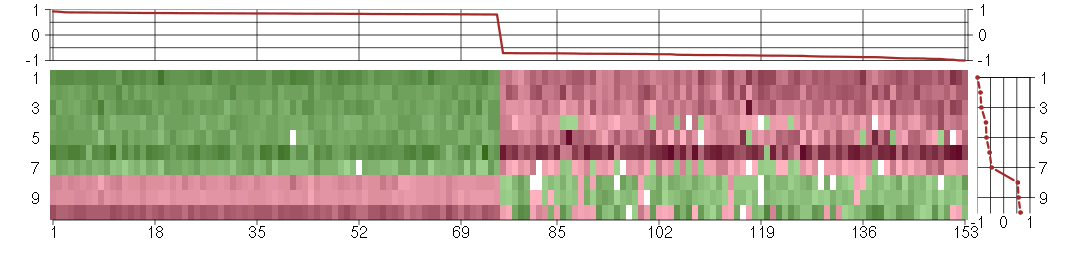

Under-expression is coded with green,
over-expression with red color.

DNA replication
The process whereby new strands of DNA are synthesized. The template for replication can either be an existing DNA molecule or RNA.
chromosome segregation
The process by which genetic material, in the form of chromosomes, is organized and then physically separated and apportioned to two or more sets.
chromosome organization
A process that is carried out at the cellular level that results in the formation, arrangement of constituent parts, or disassembly of chromosomes, structures composed of a very long molecule of DNA and associated proteins that carries hereditary information.
mitotic sister chromatid segregation
The cell cycle process whereby replicated homologous chromosomes are organized and then physically separated and apportioned to two sets during the mitotic cell cycle. Each replicated chromosome, composed of two sister chromatids, aligns at the cell equator, paired with its homologous partner. One homolog of each morphologic type goes into each of the resulting chromosome sets.
cell cycle checkpoint
The cell cycle regulatory process by which progression through the cycle can be halted until conditions are suitable for the cell to proceed to the next stage.
S phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through S phase, the part of the mitotic cell cycle during which DNA synthesis takes place.
G2 phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through G2 phase, one of two 'gap' phases in the mitotic cell cycle; G2 is the interval between the completion of DNA synthesis and the beginning of mitosis.
M phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the mitotic cell cycle during which mitosis takes place.
microtubule cytoskeleton organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of cytoskeletal structures comprising microtubules and their associated proteins.
mitotic cell cycle
Progression through the phases of the mitotic cell cycle, the most common eukaryotic cell cycle, which canonically comprises four successive phases called G1, S, G2, and M and includes replication of the genome and the subsequent segregation of chromosomes into daughter cells. In some variant cell cycles nuclear replication or nuclear division may not be followed by cell division, or G1 and G2 phases may be absent.
M phase
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the cell cycle comprising nuclear division.
nuclear division
A process by which a cell nucleus is divided into two nuclei, with DNA and other nuclear contents distributed between the daughter nuclei.
sister chromatid segregation
The process by which sister chromatids are organized and then physically separated and apportioned to two or more sets.
metabolic process
The chemical reactions and pathways, including anabolism and catabolism, by which living organisms transform chemical substances. Metabolic processes typically transform small molecules, but also include macromolecular processes such as DNA repair and replication, and protein synthesis and degradation.
regulation of protein amino acid phosphorylation
Any process that modulates the frequency, rate or extent of addition of phosphate groups into an amino acid in a protein.
positive regulation of protein amino acid phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to amino acids within a protein.
regulation of cell cycle
Any process that modulates the rate or extent of progression through the cell cycle.
cytoskeleton organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of cytoskeletal structures.
nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
The chemical reactions and pathways involving nucleobases, nucleosides, nucleotides and nucleic acids.
DNA metabolic process
The chemical reactions and pathways involving DNA, deoxyribonucleic acid, one of the two main types of nucleic acid, consisting of a long, unbranched macromolecule formed from one, or more commonly, two, strands of linked deoxyribonucleotides.
DNA-dependent DNA replication
The process whereby new strands of DNA are synthesized, using parental DNA as a template for the DNA-dependent DNA polymerases that synthesize the new strands.
DNA replication initiation
The process by which DNA replication is started; this involves the separation of a stretch of the DNA double helix, the recruitment of DNA polymerases and the initiation of polymerase action.
DNA packaging
Any process by which DNA and associated proteins are formed into a compact, orderly structure.
protein modification process
The covalent alteration of one or more amino acids occurring in proteins, peptides and nascent polypeptides (co-translational, post-translational modifications). Includes the modification of charged tRNAs that are destined to occur in a protein (pre-translation modification).
protein amino acid phosphorylation
The process of introducing a phosphate group on to a protein.
phosphorus metabolic process
The chemical reactions and pathways involving the nonmetallic element phosphorus or compounds that contain phosphorus, usually in the form of a phosphate group (PO4).
phosphate metabolic process
The chemical reactions and pathways involving the phosphate group, the anion or salt of any phosphoric acid.
organelle organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of an organelle within a cell. An organelle is an organized structure of distinctive morphology and function. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
microtubule-based process
Any cellular process that depends upon or alters the microtubule cytoskeleton, that part of the cytoskeleton comprising microtubules and their associated proteins.
cell cycle
The progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events. Canonically, the cell cycle comprises the replication and segregation of genetic material followed by the division of the cell, but in endocycles or syncytial cells nuclear replication or nuclear division may not be followed by cell division.
spindle organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of the spindle, the array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during DNA segregation and serves to move the duplicated chromosomes apart.
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
mitotic chromosome condensation
The cell cycle process whereby chromatin structure is compacted prior to mitosis in eukaryotic cells.
mitotic cell cycle checkpoint
A signal transduction-based surveillance mechanism that ensures accurate chromosome replication and segregation by preventing progression through a mitotic cell cycle until conditions are suitable for the cell to proceed to the next stage.
mitotic cell cycle spindle assembly checkpoint
A signal transduction based surveillance mechanism that ensures the fidelity of cell division by preventing the premature advance of cells from metaphase to anaphase prior to the successful attachment of kinetochores to spindle microtubules (spindle assembly).
regulation of mitotic cell cycle
Any process that modulates the rate or extent of progress through the mitotic cell cycle.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
cell proliferation
The multiplication or reproduction of cells, resulting in the expansion of a cell population.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
biosynthetic process
The chemical reactions and pathways resulting in the formation of substances; typically the energy-requiring part of metabolism in which simpler substances are transformed into more complex ones.
macromolecule biosynthetic process
The chemical reactions and pathways resulting in the formation of a macromolecule, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
positive regulation of metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways within a cell or an organism.
cellular process
Any process that is carried out at the cellular level, but not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
positive regulation of phosphorus metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving phosphorus or compounds containing phosphorus.
regulation of cell cycle process
Any process that modulates a cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
positive regulation of macromolecule metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
cellular component organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of a cellular component.
phosphorylation
The process of introducing a phosphate group into a molecule, usually with the formation of a phosphoric ester, a phosphoric anhydride or a phosphoric amide.
peptidyl-serine phosphorylation
The posttranslational phosphorylation of peptidyl-serine to form peptidyl-O-phospho-L-serine.
peptidyl-amino acid modification
The alteration of an amino acid residue in a peptide.
peptidyl-serine modification
The modification of peptidyl-serine.
regulation of phosphate metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving phosphates.
regulation of metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways within a cell or an organism.
protein metabolic process
The chemical reactions and pathways involving a specific protein, rather than of proteins in general. Includes protein modification.
cell cycle process
A cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
cell cycle phase
A cell cycle process comprising the steps by which a cell progresses through one of the biochemical and morphological phases and events that occur during successive cell replication or nuclear replication events.
chromosome condensation
The progressive compaction of dispersed interphase chromatin into threadlike chromosomes prior to mitotic or meiotic nuclear division, or during apoptosis, in eukaryotic cells.
regulation of cellular metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
positive regulation of cellular metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
regulation of protein modification process
Any process that modulates the frequency, rate or extent of the covalent alteration of one or more amino acid residues within a protein.
positive regulation of protein modification process
Any process that activates or increases the frequency, rate or extent of the covalent alteration of one or more amino acid residues within a protein.
spindle checkpoint
A cell cycle checkpoint that delays the metaphase/anaphase transition until the spindle is correctly assembled and chromosomes are attached to the spindle.
regulation of cellular protein metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
positive regulation of cellular protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
regulation of peptidyl-serine phosphorylation
Any process that modulates the frequency, rate or extent of the phosphorylation of peptidyl-serine.
positive regulation of peptidyl-serine phosphorylation
Any process that activates or increases the frequency, rate or extent of the phosphorylation of peptidyl-serine.
cellular macromolecule biosynthetic process
The chemical reactions and pathways resulting in the formation of a macromolecule, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass, carried out by individual cells.
cellular biopolymer metabolic process
The chemical reactions and pathways involving biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins, as carried out by individual cells.
cellular biopolymer biosynthetic process
The chemical reactions and pathways resulting in the formation of biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins, as carried out by individual cells.
regulation of cell proliferation
Any process that modulates the frequency, rate or extent of cell proliferation.
regulation of phosphorylation
Any process that modulates the frequency, rate or extent of addition of phosphate groups into a molecule.
positive regulation of phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to a molecule.
macromolecule metabolic process
The chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
biopolymer metabolic process
The chemical reactions and pathways involving biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins.
biopolymer biosynthetic process
The chemical reactions and pathways resulting in the formation of biopolymers, long, repeating chains of monomers found in nature e.g. polysaccharides and proteins.
biopolymer modification
The covalent alteration of one or more monomeric units in a polypeptide, polynucleotide, polysaccharide, or other biological polymer, resulting in a change in its properties.
post-translational protein modification
The covalent alteration of one or more amino acids occurring in a protein after the protein has been completely translated and released from the ribosome.
cellular metabolic process
The chemical reactions and pathways by which individual cells transform chemical substances.
primary metabolic process
The chemical reactions and pathways involving those compounds which are formed as a part of the normal anabolic and catabolic processes. These processes take place in most, if not all, cells of the organism.
cellular biosynthetic process
The chemical reactions and pathways resulting in the formation of substances, carried out by individual cells.
cellular macromolecule metabolic process
The chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass, as carried out by individual cells.
cellular protein metabolic process
The chemical reactions and pathways involving a specific protein, rather than of proteins in general, occurring at the level of an individual cell. Includes protein modification.
positive regulation of phosphate metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving phosphates.
organelle fission
The creation of two or more organelles by division of one organelle.
positive regulation of biological process
Any process that activates or increases the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of phosphorus metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving phosphorus or compounds containing phosphorus.
regulation of protein metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving a protein.
positive regulation of protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein.
cell division
The process resulting in the physical partitioning and separation of a cell into daughter cells.
G2 phase
A cell cycle process comprising the steps by which a cell progresses through G2 phase, one of two 'gap' phases in the cell cycle; G2 is the interval between the completion of DNA synthesis and the beginning of DNA segregation (usually by mitosis or meiosis).
S phase
A cell cycle process comprising the steps by which a cell progresses through S phase, the part of the cell cycle during which DNA synthesis takes place.
interphase
A cell cycle process comprising the steps by which a cell progresses through interphase, the stage of cell cycle between successive rounds of chromosome segregation. Canonically, interphase is the stage of the cell cycle during which the biochemical and physiologic functions of the cell are performed and replication of chromatin occurs.
interphase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through interphase, the stage of cell cycle between successive rounds of mitosis. Canonically, interphase is the stage of the cell cycle during which the biochemical and physiologic functions of the cell are performed and replication of chromatin occurs.
regulation of macromolecule metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
biological regulation
Any process that modulates the frequency, rate or extent of any biological process, quality or function.
all
This term is the most general term possible
cellular metabolic process
The chemical reactions and pathways by which individual cells transform chemical substances.
positive regulation of metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways within a cell or an organism.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways within a cell or an organism.
positive regulation of biological process
Any process that activates or increases the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
positive regulation of metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways within a cell or an organism.
macromolecule biosynthetic process
The chemical reactions and pathways resulting in the formation of a macromolecule, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
positive regulation of macromolecule metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
regulation of macromolecule metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
regulation of cellular metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
positive regulation of cellular metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
cellular biosynthetic process
The chemical reactions and pathways resulting in the formation of substances, carried out by individual cells.
cellular macromolecule metabolic process
The chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass, as carried out by individual cells.
nucleobase, nucleoside, nucleotide and nucleic acid metabolic process
The chemical reactions and pathways involving nucleobases, nucleosides, nucleotides and nucleic acids.
cell cycle process
A cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
sister chromatid segregation
The process by which sister chromatids are organized and then physically separated and apportioned to two or more sets.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
positive regulation of cellular metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
regulation of cell cycle
Any process that modulates the rate or extent of progression through the cell cycle.
regulation of cellular metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
regulation of cell proliferation
Any process that modulates the frequency, rate or extent of cell proliferation.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
cellular macromolecule biosynthetic process
The chemical reactions and pathways resulting in the formation of a macromolecule, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass, carried out by individual cells.
positive regulation of cellular metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways by which individual cells transform chemical substances.
positive regulation of macromolecule metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving macromolecules, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass.
protein metabolic process
The chemical reactions and pathways involving a specific protein, rather than of proteins in general. Includes protein modification.
biopolymer biosynthetic process
The chemical reactions and pathways resulting in the formation of biopolymers, long, repeating chains of monomers found in nature e.g. polysaccharides and proteins.
cellular macromolecule biosynthetic process
The chemical reactions and pathways resulting in the formation of a macromolecule, any molecule of high relative molecular mass, the structure of which essentially comprises the multiple repetition of units derived, actually or conceptually, from molecules of low relative molecular mass, carried out by individual cells.
cellular biopolymer metabolic process
The chemical reactions and pathways involving biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins, as carried out by individual cells.
positive regulation of phosphorus metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving phosphorus or compounds containing phosphorus.
regulation of phosphorus metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving phosphorus or compounds containing phosphorus.
regulation of protein metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving a protein.
positive regulation of protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein.
regulation of mitotic cell cycle
Any process that modulates the rate or extent of progress through the mitotic cell cycle.
regulation of cell cycle process
Any process that modulates a cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
cellular biopolymer biosynthetic process
The chemical reactions and pathways resulting in the formation of biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins, as carried out by individual cells.
positive regulation of cellular protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
positive regulation of cellular protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
positive regulation of phosphorus metabolic process
Any process that increases the frequency, rate or extent of the chemical reactions and pathways involving phosphorus or compounds containing phosphorus.
regulation of cellular protein metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
positive regulation of protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein.
DNA metabolic process
The chemical reactions and pathways involving DNA, deoxyribonucleic acid, one of the two main types of nucleic acid, consisting of a long, unbranched macromolecule formed from one, or more commonly, two, strands of linked deoxyribonucleotides.
cellular biopolymer biosynthetic process
The chemical reactions and pathways resulting in the formation of biopolymers, long, repeating chains of monomers found in nature, such as polysaccharides and proteins, as carried out by individual cells.
cellular protein metabolic process
The chemical reactions and pathways involving a specific protein, rather than of proteins in general, occurring at the level of an individual cell. Includes protein modification.
regulation of phosphate metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving phosphates.
positive regulation of phosphate metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving phosphates.
protein modification process
The covalent alteration of one or more amino acids occurring in proteins, peptides and nascent polypeptides (co-translational, post-translational modifications). Includes the modification of charged tRNAs that are destined to occur in a protein (pre-translation modification).
regulation of cellular protein metabolic process
Any process that modulates the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
positive regulation of cellular protein metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving a protein, occurring at the level of an individual cell.
mitotic chromosome condensation
The cell cycle process whereby chromatin structure is compacted prior to mitosis in eukaryotic cells.
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
mitotic cell cycle checkpoint
A signal transduction-based surveillance mechanism that ensures accurate chromosome replication and segregation by preventing progression through a mitotic cell cycle until conditions are suitable for the cell to proceed to the next stage.
mitotic chromosome condensation
The cell cycle process whereby chromatin structure is compacted prior to mitosis in eukaryotic cells.
sister chromatid segregation
The process by which sister chromatids are organized and then physically separated and apportioned to two or more sets.
chromosome condensation
The progressive compaction of dispersed interphase chromatin into threadlike chromosomes prior to mitotic or meiotic nuclear division, or during apoptosis, in eukaryotic cells.
microtubule cytoskeleton organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of cytoskeletal structures comprising microtubules and their associated proteins.
M phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the mitotic cell cycle during which mitosis takes place.
spindle organization
A process that is carried out at the cellular level which results in the formation, arrangement of constituent parts, or disassembly of the spindle, the array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during DNA segregation and serves to move the duplicated chromosomes apart.
mitotic sister chromatid segregation
The cell cycle process whereby replicated homologous chromosomes are organized and then physically separated and apportioned to two sets during the mitotic cell cycle. Each replicated chromosome, composed of two sister chromatids, aligns at the cell equator, paired with its homologous partner. One homolog of each morphologic type goes into each of the resulting chromosome sets.
G2 phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through G2 phase, one of two 'gap' phases in the mitotic cell cycle; G2 is the interval between the completion of DNA synthesis and the beginning of mitosis.
S phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through S phase, the part of the mitotic cell cycle during which DNA synthesis takes place.
G2 phase
A cell cycle process comprising the steps by which a cell progresses through G2 phase, one of two 'gap' phases in the cell cycle; G2 is the interval between the completion of DNA synthesis and the beginning of DNA segregation (usually by mitosis or meiosis).
S phase
A cell cycle process comprising the steps by which a cell progresses through S phase, the part of the cell cycle during which DNA synthesis takes place.
interphase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through interphase, the stage of cell cycle between successive rounds of mitosis. Canonically, interphase is the stage of the cell cycle during which the biochemical and physiologic functions of the cell are performed and replication of chromatin occurs.
DNA replication
The process whereby new strands of DNA are synthesized. The template for replication can either be an existing DNA molecule or RNA.
positive regulation of protein modification process
Any process that activates or increases the frequency, rate or extent of the covalent alteration of one or more amino acid residues within a protein.
positive regulation of phosphate metabolic process
Any process that activates or increases the frequency, rate or extent of the chemical reactions and pathways involving phosphates.
regulation of protein modification process
Any process that modulates the frequency, rate or extent of the covalent alteration of one or more amino acid residues within a protein.
positive regulation of protein modification process
Any process that activates or increases the frequency, rate or extent of the covalent alteration of one or more amino acid residues within a protein.
regulation of phosphorylation
Any process that modulates the frequency, rate or extent of addition of phosphate groups into a molecule.
positive regulation of phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to a molecule.
mitotic cell cycle spindle assembly checkpoint
A signal transduction based surveillance mechanism that ensures the fidelity of cell division by preventing the premature advance of cells from metaphase to anaphase prior to the successful attachment of kinetochores to spindle microtubules (spindle assembly).
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
positive regulation of protein amino acid phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to amino acids within a protein.
positive regulation of protein amino acid phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to amino acids within a protein.
regulation of protein amino acid phosphorylation
Any process that modulates the frequency, rate or extent of addition of phosphate groups into an amino acid in a protein.
positive regulation of phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to a molecule.
protein amino acid phosphorylation
The process of introducing a phosphate group on to a protein.
DNA replication initiation
The process by which DNA replication is started; this involves the separation of a stretch of the DNA double helix, the recruitment of DNA polymerases and the initiation of polymerase action.
regulation of protein amino acid phosphorylation
Any process that modulates the frequency, rate or extent of addition of phosphate groups into an amino acid in a protein.
positive regulation of protein amino acid phosphorylation
Any process that activates or increases the frequency, rate or extent of addition of phosphate groups to amino acids within a protein.
positive regulation of peptidyl-serine phosphorylation
Any process that activates or increases the frequency, rate or extent of the phosphorylation of peptidyl-serine.
peptidyl-serine phosphorylation
The posttranslational phosphorylation of peptidyl-serine to form peptidyl-O-phospho-L-serine.
regulation of peptidyl-serine phosphorylation
Any process that modulates the frequency, rate or extent of the phosphorylation of peptidyl-serine.
positive regulation of peptidyl-serine phosphorylation
Any process that activates or increases the frequency, rate or extent of the phosphorylation of peptidyl-serine.

nuclear chromosome
A chromosome found in the nucleus of a eukaryotic cell.
condensed chromosome
A highly compacted molecule of DNA and associated proteins resulting in a cytologically distinct structure.
intracellular
The living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
chromosome, centromeric region
The region of a chromosome that includes the centromere and associated proteins. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
condensed chromosome kinetochore
A multisubunit complex that is located at the centromeric region of a condensed chromosome and provides an attachment point for the spindle microtubules.
condensed chromosome, centromeric region
The region of a condensed chromosome that includes the centromere and associated proteins, including the kinetochore. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
chromatin
The ordered and organized complex of DNA and protein that forms the chromosome.
condensin complex
A multisubunit protein complex that plays a central role in chromosome condensation.
spindle pole
Either of the ends of a spindle, where spindle microtubules are organized; usually contains a microtubule organizing center and accessory molecules, spindle microtubules and astral microtubules.
extracellular region
The space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers the host cell environment outside an intracellular parasite.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
extracellular space
That part of a multicellular organism outside the cells proper, usually taken to be outside the plasma membranes, and occupied by fluid.
cell
The basic structural and functional unit of all organisms. Includes the plasma membrane and any external encapsulating structures such as the cell wall and cell envelope.
nucleus
A membrane-bounded organelle of eukaryotic cells in which chromosomes are housed and replicated. In most cells, the nucleus contains all of the cell's chromosomes except the organellar chromosomes, and is the site of RNA synthesis and processing. In some species, or in specialized cell types, RNA metabolism or DNA replication may be absent.
replication fork
The Y-shaped region of a replicating DNA molecule, resulting from the separation of the DNA strands and in which the synthesis of new strands takes place. Also includes associated protein complexes.
chromosome
A structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
cytoskeleton
Any of the various filamentous elements that form the internal framework of cells, and typically remain after treatment of the cells with mild detergent to remove membrane constituents and soluble components of the cytoplasm. The term embraces intermediate filaments, microfilaments, microtubules, the microtrabecular lattice, and other structures characterized by a polymeric filamentous nature and long-range order within the cell. The various elements of the cytoskeleton not only serve in the maintenance of cellular shape but also have roles in other cellular functions, including cellular movement, cell division, endocytosis, and movement of organelles.
microtubule
Any of the long, generally straight, hollow tubes of internal diameter 12-15 nm and external diameter 24 nm found in a wide variety of eukaryotic cells; each consists (usually) of 13 protofilaments of polymeric tubulin, staggered in such a manner that the tubulin monomers are arranged in a helical pattern on the microtubular surface, and with the alpha/beta axes of the tubulin subunits parallel to the long axis of the tubule; exist in equilibrium with pool of tubulin monomers and can be rapidly assembled or disassembled in response to physiological stimuli; concerned with force generation, e.g. in the spindle.
microtubule cytoskeleton
The part of the cytoskeleton (the internal framework of a cell) composed of microtubules and associated proteins.
macromolecular complex
A stable assembly of two or more macromolecules, i.e. proteins, nucleic acids, carbohydrates or lipids, in which the constituent parts function together.
organelle
Organized structure of distinctive morphology and function. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
membrane-bounded organelle
Organized structure of distinctive morphology and function, bounded by a single or double lipid bilayer membrane. Includes the nucleus, mitochondria, plastids, vacuoles, and vesicles. Excludes the plasma membrane.
non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane. Includes ribosomes, the cytoskeleton and chromosomes.
intracellular organelle
Organized structure of distinctive morphology and function, occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
intracellular membrane-bounded organelle
Organized structure of distinctive morphology and function, bounded by a single or double lipid bilayer membrane and occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, and vesicles. Excludes the plasma membrane.
intracellular non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane and occurring within the cell. Includes ribosomes, the cytoskeleton and chromosomes.
protein complex
Any macromolecular complex composed of two or more polypeptide subunits, which may or may not be identical. Protein complexes may have other associated non-protein prosthetic groups, such as nucleotides, metal ions or carbohydrate groups.
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.
organelle part
Any constituent part of an organelle, an organized structure of distinctive morphology and function. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton, but excludes the plasma membrane.
intracellular part
Any constituent part of the living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
chromosomal part
Any constituent part of a chromosome, a structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
nuclear part
Any constituent part of the nucleus, a membrane-bounded organelle of eukaryotic cells in which chromosomes are housed and replicated.
cytoskeletal part
Any constituent part of the cytoskeleton, a cellular scaffolding or skeleton that maintains cell shape, enables some cell motion (using structures such as flagella and cilia), and plays important roles in both intra-cellular transport (e.g. the movement of vesicles and organelles) and cellular division. Includes constituent parts of intermediate filaments, microfilaments, microtubules, and the microtrabecular lattice.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
nuclear chromosome part
Any constituent part of a nuclear chromosome, a chromosome found in the nucleus of a eukaryotic cell.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
all
This term is the most general term possible
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
organelle part
Any constituent part of an organelle, an organized structure of distinctive morphology and function. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton, but excludes the plasma membrane.
intracellular membrane-bounded organelle
Organized structure of distinctive morphology and function, bounded by a single or double lipid bilayer membrane and occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, and vesicles. Excludes the plasma membrane.
intracellular non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane and occurring within the cell. Includes ribosomes, the cytoskeleton and chromosomes.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
intracellular part
Any constituent part of the living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
intracellular organelle
Organized structure of distinctive morphology and function, occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
condensin complex
A multisubunit protein complex that plays a central role in chromosome condensation.
nuclear chromosome part
Any constituent part of a nuclear chromosome, a chromosome found in the nucleus of a eukaryotic cell.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
nuclear part
Any constituent part of the nucleus, a membrane-bounded organelle of eukaryotic cells in which chromosomes are housed and replicated.
nuclear chromosome
A chromosome found in the nucleus of a eukaryotic cell.
chromosomal part
Any constituent part of a chromosome, a structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
spindle pole
Either of the ends of a spindle, where spindle microtubules are organized; usually contains a microtubule organizing center and accessory molecules, spindle microtubules and astral microtubules.
cytoskeletal part
Any constituent part of the cytoskeleton, a cellular scaffolding or skeleton that maintains cell shape, enables some cell motion (using structures such as flagella and cilia), and plays important roles in both intra-cellular transport (e.g. the movement of vesicles and organelles) and cellular division. Includes constituent parts of intermediate filaments, microfilaments, microtubules, and the microtrabecular lattice.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
nuclear chromosome part
Any constituent part of a nuclear chromosome, a chromosome found in the nucleus of a eukaryotic cell.
condensed chromosome, centromeric region
The region of a condensed chromosome that includes the centromere and associated proteins, including the kinetochore. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
condensin complex
A multisubunit protein complex that plays a central role in chromosome condensation.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
microtubule
Any of the long, generally straight, hollow tubes of internal diameter 12-15 nm and external diameter 24 nm found in a wide variety of eukaryotic cells; each consists (usually) of 13 protofilaments of polymeric tubulin, staggered in such a manner that the tubulin monomers are arranged in a helical pattern on the microtubular surface, and with the alpha/beta axes of the tubulin subunits parallel to the long axis of the tubule; exist in equilibrium with pool of tubulin monomers and can be rapidly assembled or disassembled in response to physiological stimuli; concerned with force generation, e.g. in the spindle.
condensed chromosome kinetochore
A multisubunit complex that is located at the centromeric region of a condensed chromosome and provides an attachment point for the spindle microtubules.

protein binding
Interacting selectively with any protein or protein complex (a complex of two or more proteins that may include other nonprotein molecules).
molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
motor activity
Catalysis of movement along a polymeric molecule such as a microfilament or microtubule, coupled to the hydrolysis of a nucleoside triphosphate.
microtubule motor activity
Catalysis of movement along a microtubule, coupled to the hydrolysis of a nucleoside triphosphate (usually ATP).
catalytic activity
Catalysis of a biochemical reaction at physiological temperatures. In biologically catalyzed reactions, the reactants are known as substrates, and the catalysts are naturally occurring macromolecular substances known as enzymes. Enzymes possess specific binding sites for substrates, and are usually composed wholly or largely of protein, but RNA that has catalytic activity (ribozyme) is often also regarded as enzymatic.
nucleoside-triphosphatase activity
Catalysis of the reaction: a nucleoside triphosphate + H2O = nucleoside diphosphate + phosphate.
receptor binding
Interacting selectively with one or more specific sites on a receptor molecule, a macromolecule that undergoes combination with a hormone, neurotransmitter, drug or intracellular messenger to initiate a change in cell function.
cytokine activity
Functions to control the survival, growth, differentiation and effector function of tissues and cells.
binding
The selective, often stoichiometric, interaction of a molecule with one or more specific sites on another molecule.
growth factor activity
The function that stimulates a cell to grow or proliferate. Most growth factors have other actions besides the induction of cell growth or proliferation.
pyrophosphatase activity
Catalysis of the hydrolysis of a pyrophosphate bond between two phosphate groups, leaving one phosphate on each of the two fragments.
hydrolase activity
Catalysis of the hydrolysis of various bonds, e.g. C-O, C-N, C-C, phosphoric anhydride bonds, etc. Hydrolase is the systematic name for any enzyme of EC class 3.
hydrolase activity, acting on acid anhydrides
Catalysis of the hydrolysis of any acid anhydride.
hydrolase activity, acting on acid anhydrides, in phosphorus-containing anhydrides
Catalysis of the hydrolysis of any acid anhydride which contains phosphorus.
all
This term is the most general term possible
| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 03030 | 1.209e-04 | 0.7734 | 8 | 33 | DNA replication |
| 04110 | 1.618e-03 | 2.297 | 11 | 98 | Cell cycle |
AJAP1adherens junctions associated protein 1 (206460_at), score: -0.76 ANGPTL4angiopoietin-like 4 (221009_s_at), score: -0.81 AREGamphiregulin (205239_at), score: -0.8 ASF1BASF1 anti-silencing function 1 homolog B (S. cerevisiae) (218115_at), score: 0.83 ASPMasp (abnormal spindle) homolog, microcephaly associated (Drosophila) (219918_s_at), score: 0.84 ATP13A3ATPase type 13A3 (219558_at), score: -0.8 AURKBaurora kinase B (209464_at), score: 0.91 B3GNT2UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2 (219326_s_at), score: -0.81 BIRC5baculoviral IAP repeat-containing 5 (202095_s_at), score: 0.89 BLMBloom syndrome (205733_at), score: 0.85 BMP2bone morphogenetic protein 2 (205289_at), score: -0.87 BMP6bone morphogenetic protein 6 (206176_at), score: -0.94 BRCA2breast cancer 2, early onset (208368_s_at), score: 0.85 BUB1budding uninhibited by benzimidazoles 1 homolog (yeast) (209642_at), score: 0.85 BUB1Bbudding uninhibited by benzimidazoles 1 homolog beta (yeast) (203755_at), score: 0.83 C17orf91chromosome 17 open reading frame 91 (214696_at), score: -1 C18orf24chromosome 18 open reading frame 24 (217640_x_at), score: 0.86 C21orf45chromosome 21 open reading frame 45 (219004_s_at), score: 0.81 CBLBCas-Br-M (murine) ecotropic retroviral transforming sequence b (209682_at), score: -0.78 CBR3carbonyl reductase 3 (205379_at), score: 0.88 CCNB2cyclin B2 (202705_at), score: 0.86 CCNJcyclin J (219470_x_at), score: -0.72 CDC20cell division cycle 20 homolog (S. cerevisiae) (202870_s_at), score: 0.85 CDC25Bcell division cycle 25 homolog B (S. pombe) (201853_s_at), score: 0.8 CDC45LCDC45 cell division cycle 45-like (S. cerevisiae) (204126_s_at), score: 0.81 CDCA3cell division cycle associated 3 (221436_s_at), score: 0.85 CDCA8cell division cycle associated 8 (221520_s_at), score: 0.83 CDT1chromatin licensing and DNA replication factor 1 (209832_s_at), score: 0.82 CDYLchromodomain protein, Y-like (203100_s_at), score: -0.72 CENPFcentromere protein F, 350/400ka (mitosin) (207828_s_at), score: 0.86 CENPMcentromere protein M (218741_at), score: 0.88 CEP55centrosomal protein 55kDa (218542_at), score: 0.84 CHAF1Achromatin assembly factor 1, subunit A (p150) (214426_x_at), score: 0.83 CHMP1Bchromatin modifying protein 1B (218178_s_at), score: -0.79 CKAP2cytoskeleton associated protein 2 (218252_at), score: 0.87 COL14A1collagen, type XIV, alpha 1 (212865_s_at), score: -0.73 COL15A1collagen, type XV, alpha 1 (203477_at), score: -0.72 CPA3carboxypeptidase A3 (mast cell) (205624_at), score: -0.75 CSGALNACT2chondroitin sulfate N-acetylgalactosaminyltransferase 2 (222235_s_at), score: -0.75 CXCR7chemokine (C-X-C motif) receptor 7 (212977_at), score: -0.74 DDX11DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 11 (CHL1-like helicase homolog, S. cerevisiae) (208149_x_at), score: 0.85 DEPDC1DEP domain containing 1 (220295_x_at), score: 0.87 DLGAP5discs, large (Drosophila) homolog-associated protein 5 (203764_at), score: 0.84 ENTPD7ectonucleoside triphosphate diphosphohydrolase 7 (220153_at), score: -1 F10coagulation factor X (205620_at), score: -0.74 FABP3fatty acid binding protein 3, muscle and heart (mammary-derived growth inhibitor) (214285_at), score: -0.79 FAM64Afamily with sequence similarity 64, member A (221591_s_at), score: 0.85 FANCIFanconi anemia, complementation group I (213007_at), score: 0.82 FEM1Bfem-1 homolog b (C. elegans) (212367_at), score: -0.71 FGF1fibroblast growth factor 1 (acidic) (205117_at), score: -0.84 FGF7fibroblast growth factor 7 (keratinocyte growth factor) (205782_at), score: -0.94 FOXM1forkhead box M1 (202580_x_at), score: 0.83 GABARAPL1GABA(A) receptor-associated protein like 1 (208868_s_at), score: -0.74 GEMGTP binding protein overexpressed in skeletal muscle (204472_at), score: -0.81 GFPT2glutamine-fructose-6-phosphate transaminase 2 (205100_at), score: -0.86 GINS2GINS complex subunit 2 (Psf2 homolog) (221521_s_at), score: 0.86 GPR183G protein-coupled receptor 183 (205419_at), score: -0.97 GTSE1G-2 and S-phase expressed 1 (204318_s_at), score: 0.85 HBEGFheparin-binding EGF-like growth factor (203821_at), score: -0.88 HJURPHolliday junction recognition protein (218726_at), score: 0.82 HK2hexokinase 2 (202934_at), score: -0.74 HMGB2high-mobility group box 2 (208808_s_at), score: 0.86 HMMRhyaluronan-mediated motility receptor (RHAMM) (207165_at), score: 0.85 HTR2A5-hydroxytryptamine (serotonin) receptor 2A (207135_at), score: -0.9 ICAM1intercellular adhesion molecule 1 (202638_s_at), score: -0.86 IDI2isopentenyl-diphosphate delta isomerase 2 (217631_at), score: -0.72 IL11interleukin 11 (206924_at), score: -0.87 IL6interleukin 6 (interferon, beta 2) (205207_at), score: -0.92 IL8interleukin 8 (211506_s_at), score: -0.85 INSIG1insulin induced gene 1 (201627_s_at), score: -0.74 ITGB3BPintegrin beta 3 binding protein (beta3-endonexin) (205176_s_at), score: 0.81 JHDM1Djumonji C domain containing histone demethylase 1 homolog D (S. cerevisiae) (221778_at), score: -0.82 KIAA0586KIAA0586 (205631_at), score: 0.87 KIAA1644KIAA1644 (52837_at), score: -0.8 KIF11kinesin family member 11 (204444_at), score: 0.84 KIF15kinesin family member 15 (219306_at), score: 0.85 KIF18Bkinesin family member 18B (222039_at), score: 0.87 KIF20Bkinesin family member 20B (205235_s_at), score: 0.82 KIF22kinesin family member 22 (202183_s_at), score: 0.88 KIF4Akinesin family member 4A (218355_at), score: 0.89 KLHL21kelch-like 21 (Drosophila) (203068_at), score: -0.79 LIFleukemia inhibitory factor (cholinergic differentiation factor) (205266_at), score: -0.82 LMCD1LIM and cysteine-rich domains 1 (218574_s_at), score: -0.77 LMNB1lamin B1 (203276_at), score: 0.82 MAD2L1MAD2 mitotic arrest deficient-like 1 (yeast) (203362_s_at), score: 0.83 MCM10minichromosome maintenance complex component 10 (220651_s_at), score: 0.83 MCM2minichromosome maintenance complex component 2 (202107_s_at), score: 0.85 MCM5minichromosome maintenance complex component 5 (216237_s_at), score: 0.92 MCM7minichromosome maintenance complex component 7 (210983_s_at), score: 0.89 MKI67antigen identified by monoclonal antibody Ki-67 (212022_s_at), score: 0.82 MLF1IPMLF1 interacting protein (218883_s_at), score: 0.82 NCAPD2non-SMC condensin I complex, subunit D2 (201774_s_at), score: 0.82 NCAPGnon-SMC condensin I complex, subunit G (218663_at), score: 0.83 NCAPG2non-SMC condensin II complex, subunit G2 (219588_s_at), score: 0.8 NCAPHnon-SMC condensin I complex, subunit H (212949_at), score: 0.82 NDC80NDC80 homolog, kinetochore complex component (S. cerevisiae) (204162_at), score: 0.84 NDEL1nudE nuclear distribution gene E homolog (A. nidulans)-like 1 (208093_s_at), score: -0.75 NEIL3nei endonuclease VIII-like 3 (E. coli) (219502_at), score: 0.81 NEK2NIMA (never in mitosis gene a)-related kinase 2 (204641_at), score: 0.82 NFATC1nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (210162_s_at), score: -0.72 NR3C1nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) (201866_s_at), score: -0.79 NR4A1nuclear receptor subfamily 4, group A, member 1 (202340_x_at), score: -0.72 NR4A3nuclear receptor subfamily 4, group A, member 3 (209959_at), score: -0.87 NUPL1nucleoporin like 1 (204435_at), score: -0.72 NUSAP1nucleolar and spindle associated protein 1 (218039_at), score: 0.85 OIP5Opa interacting protein 5 (213599_at), score: 0.82 PCTK2PCTAIRE protein kinase 2 (221918_at), score: -0.71 PDGFDplatelet derived growth factor D (219304_s_at), score: -0.72 PIK3CDphosphoinositide-3-kinase, catalytic, delta polypeptide (203879_at), score: -0.92 PMEPA1prostate transmembrane protein, androgen induced 1 (217875_s_at), score: -0.91 POLA1polymerase (DNA directed), alpha 1, catalytic subunit (204835_at), score: 0.88 POLD1polymerase (DNA directed), delta 1, catalytic subunit 125kDa (203422_at), score: 0.87 POLEpolymerase (DNA directed), epsilon (216026_s_at), score: 0.82 PPARDperoxisome proliferator-activated receptor delta (37152_at), score: -0.82 PRIM1primase, DNA, polypeptide 1 (49kDa) (205053_at), score: 0.81 PTHLHparathyroid hormone-like hormone (211756_at), score: -0.93 PTTG1pituitary tumor-transforming 1 (203554_x_at), score: 0.84 RAD51AP1RAD51 associated protein 1 (204146_at), score: 0.81 RASGRP1RAS guanyl releasing protein 1 (calcium and DAG-regulated) (205590_at), score: -0.79 RNASEH2Aribonuclease H2, subunit A (203022_at), score: 0.86 RUNX1runt-related transcription factor 1 (209360_s_at), score: -0.83 SGK1serum/glucocorticoid regulated kinase 1 (201739_at), score: -0.72 SH3TC2SH3 domain and tetratricopeptide repeats 2 (219710_at), score: 0.81 SKILSKI-like oncogene (206675_s_at), score: -0.8 SLC19A2solute carrier family 19 (thiamine transporter), member 2 (209681_at), score: -0.96 SMC2structural maintenance of chromosomes 2 (204240_s_at), score: 0.83 SOX4SRY (sex determining region Y)-box 4 (201417_at), score: -0.74 SOX9SRY (sex determining region Y)-box 9 (202935_s_at), score: -0.84 SPATA2Lspermatogenesis associated 2-like (214965_at), score: -0.85 SPRY2sprouty homolog 2 (Drosophila) (204011_at), score: -0.71 SRFserum response factor (c-fos serum response element-binding transcription factor) (202401_s_at), score: -0.81 ST3GAL1ST3 beta-galactoside alpha-2,3-sialyltransferase 1 (208322_s_at), score: -0.86 STK38Lserine/threonine kinase 38 like (212572_at), score: -0.92 SVEP1sushi, von Willebrand factor type A, EGF and pentraxin domain containing 1 (213247_at), score: -0.74 TACC3transforming, acidic coiled-coil containing protein 3 (218308_at), score: 0.86 TACSTD2tumor-associated calcium signal transducer 2 (202286_s_at), score: -0.73 TGFBR1transforming growth factor, beta receptor 1 (206943_at), score: -0.89 THBDthrombomodulin (203887_s_at), score: -0.79 TIMELESStimeless homolog (Drosophila) (203046_s_at), score: 0.86 TMEM194Atransmembrane protein 194A (212621_at), score: 0.84 TMEM39Atransmembrane protein 39A (218615_s_at), score: -0.79 TMEM41Btransmembrane protein 41B (212623_at), score: -0.74 TP53BP2tumor protein p53 binding protein, 2 (203120_at), score: -0.73 TPX2TPX2, microtubule-associated, homolog (Xenopus laevis) (210052_s_at), score: 0.88 TRIB1tribbles homolog 1 (Drosophila) (202241_at), score: -0.75 TRIP13thyroid hormone receptor interactor 13 (204033_at), score: 0.82 TRPC6transient receptor potential cation channel, subfamily C, member 6 (217287_s_at), score: -0.74 TTC17tetratricopeptide repeat domain 17 (218972_at), score: -0.81 TTKTTK protein kinase (204822_at), score: 0.84 VCLvinculin (200930_s_at), score: -0.85 VEGFAvascular endothelial growth factor A (211527_x_at), score: -0.78 YRDCyrdC domain containing (E. coli) (218647_s_at), score: -0.92 ZFP36L2zinc finger protein 36, C3H type-like 2 (201367_s_at), score: 0.84
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485851.cel | 11 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486231.cel | 30 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486351.cel | 36 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485811.cel | 9 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486331.cel | 35 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485891.cel | 13 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 6Twin.CEL | 6 | 2 | DS-twin | hgu133plus2 | none | DS-twin 6 |
| 5CTwin.CEL | 5 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 5 |
| 1Twin.CEL | 1 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 1 |