Previous module |
Next module
Module #531, TG: 3, TC: 2, 81 probes, 81 Entrez genes, 5 conditions


HELP
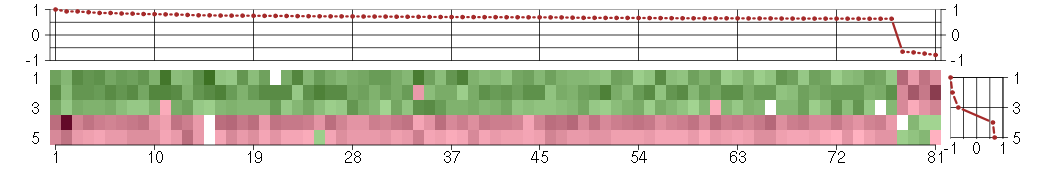
The image plot shows the color-coded level of gene expression, for the
genes and conditions in a given transcription module. The genes are on
the horizontal, the conditions on the vertical axis.
The genes are ordered according to their ISA gene scores, similarly
the conditions are ordered according to their condition scores. The
score of a gene means the «degree of inclusion» in
the module: a high score gene is essential in the module.
Condition scores can also be negative, that means that the genes of
the module are all down-regulated in the condition. Here the absolute
value of the score gives the «degree of inclusion».
The plots above and beside the expression matrix show the gene scores
and condition scores, respectively.
Note that the plot is interactive, you can see the name of the gene
and condition under the mouse cursor.
The expression matrix was normalized to have mean zero and standard
deviation one for every gene separately across all conditions
(i.e. not just for the conditions in the module).
— Click on the Help button again to close this help window.

Under-expression is coded with green,
over-expression with red color.
Help |
Hide |
Top
Help |
Show |
Top
The GO tree — Biological processes
HELP
This is one of three sections showing Gene Ontology enrichment of the
current module: in this case for biological processes.
The graph shows the hierarchy of the GO categories, their enrichment
for the current module is color coded, and the blue number beside the
category is the minus log ten p-value of the enrichment. (Calculated
using the standard hypergeometric test.) The color of the arrows code
«is a» (cyan) and «part of» relationships.
The tree was built the following way. First all GO terms with more
significant enrichment p-value than 0.05 were collected. Then all
paths from these terms to the root node of the GO tree were included
too. If a GO term is included more than once in the tree, then the
green numbers show 1) the id of the node, this makes it easier to find
other appereances of the term, and 2) the number of appearences.
Note that the same GO category might show up on the graph many
times. This is because the GO was «straightened» for this
graph, i.e. if there are more paths from a GO term to the root node of
the tree, all of them are included. The green numbers
Move the mouse cursor over the terms to get their definition. Clicking
on them takes you to the corresponding Gene Ontology web page.
If you cannot see a graph here at all, that means that there were no
significantly enriched GO categories, at the 0.05 level.
— Click on the Help button again to close this help window.

cytokine production
The appearance of a cytokine due to biosynthesis or secretion following a cellular stimulus, resulting in an increase in its intracellular or extracellular levels.
regulation of cytokine production
Any process that modulates the frequency, rate, or extent of production of a cytokine.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
multicellular organismal process
Any biological process, occurring at the level of a multicellular organism, pertinent to its function.
tumor necrosis factor production
The appearance of tumor necrosis factor due to biosynthesis or secretion following a cellular stimulus, resulting in an increase in its intracellular or extracellular levels.
regulation of tumor necrosis factor production
Any process that modulates the frequency, rate, or extent of tumor necrosis factor production.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of multicellular organismal process
Any process that modulates the frequency, rate or extent of a multicellular organismal process, the processes pertinent to the function of a multicellular organism above the cellular level; includes the integrated processes of tissues and organs.
biological regulation
Any process that modulates the frequency, rate or extent of any biological process, quality or function.
all
This term is the most general term possible
regulation of multicellular organismal process
Any process that modulates the frequency, rate or extent of a multicellular organismal process, the processes pertinent to the function of a multicellular organism above the cellular level; includes the integrated processes of tissues and organs.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of cytokine production
Any process that modulates the frequency, rate, or extent of production of a cytokine.
regulation of tumor necrosis factor production
Any process that modulates the frequency, rate, or extent of tumor necrosis factor production.
Help |
Hide |
Top
Help |
Show |
Top
The GO tree — Cellular Components
HELP
This is one of three sections showing Gene Ontology enrichment of the
current module: in this case for cellular components.
The graph shows the hierarchy of the GO categories, their enrichment
for the current module is color coded, and the blue number beside the
category is the minus log ten p-value of the enrichment. (Calculated
using the standard hypergeometric test.) The color of the arrows code
«is a» (cyan) and «part of» relationships.
The tree was built the following way. First all GO terms with more
significant enrichment p-value than 0.05 were collected. Then all
paths from these terms to the root node of the GO tree were included
too. If a GO term is included more than once in the tree, then the
green numbers show 1) the id of the node, this makes it easier to find
other appereances of the term, and 2) the number of appearences.
Note that the same GO category might show up on the graph many
times. This is because the GO was «straightened» for this
graph, i.e. if there are more paths from a GO term to the root node of
the tree, all of them are included. The green numbers
Move the mouse cursor over the terms to get their definition. Clicking
on them takes you to the corresponding Gene Ontology web page.
If you cannot see a graph here at all, that means that there were no
significantly enriched GO categories, at the 0.05 level.
— Click on the Help button again to close this help window.

Help |
Hide |
Top
Help |
Show |
Top
The GO tree — Molecular Function
HELP
This is one of three sections showing Gene Ontology enrichment of the
current module: in this case for molecular function.
The graph shows the hierarchy of the GO categories, their enrichment
for the current module is color coded, and the blue number beside the
category is the minus log ten p-value of the enrichment. (Calculated
using the standard hypergeometric test.) The color of the arrows code
«is a» (cyan) and «part of» relationships.
The tree was built the following way. First all GO terms with more
significant enrichment p-value than 0.05 were collected. Then all
paths from these terms to the root node of the GO tree were included
too. If a GO term is included more than once in the tree, then the
green numbers show 1) the id of the node, this makes it easier to find
other appereances of the term, and 2) the number of appearences.
Note that the same GO category might show up on the graph many
times. This is because the GO was «straightened» for this
graph, i.e. if there are more paths from a GO term to the root node of
the tree, all of them are included. The green numbers
Move the mouse cursor over the terms to get their definition. Clicking
on them takes you to the corresponding Gene Ontology web page.
If you cannot see a graph here at all, that means that there were no
significantly enriched GO categories, at the 0.05 level.
— Click on the Help button again to close this help window.

Help |
Hide |
Top
Help |
Show |
Top
GO BP test for over-representation
HELP
List of all enriched GO categories (biological processes), at the 0.05
p-value level.
The columns:
- ExpCount is the expected count of genes in the
module annotated with the given GO term, just by chance.
- Count
is the number of genes in the module annotated with the given GO
term.
- Size is the total number of genes (in our universe)
annotated with the GO term.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
GO category.
Clicking on the GO identifiers takes you to the Gene Ontology web
pages.
— Click on the Help button again to close this help window.
| Id | Pvalue | ExpCount | Count | Size | Term |
| GO:0032680 | 4.771e-03 | 0.07401 | 3
FCER1G, SASH3, TNFRSF8 | 9 | regulation of tumor necrosis factor production |
| GO:0032640 | 6.359e-03 | 0.08224 | 3
FCER1G, SASH3, TNFRSF8 | 10 | tumor necrosis factor production |
| GO:0032733 | 1.593e-02 | 0.02467 | 2
FCER1G, SASH3 | 3 | positive regulation of interleukin-10 production |
| GO:0032760 | 1.593e-02 | 0.02467 | 2
FCER1G, SASH3 | 3 | positive regulation of tumor necrosis factor production |
| GO:0002478 | 2.665e-02 | 0.03289 | 2
CTSE, FCER1G | 4 | antigen processing and presentation of exogenous peptide antigen |
| GO:0002495 | 2.665e-02 | 0.03289 | 2
CTSE, FCER1G | 4 | antigen processing and presentation of peptide antigen via MHC class II |
| GO:0019886 | 2.665e-02 | 0.03289 | 2
CTSE, FCER1G | 4 | antigen processing and presentation of exogenous peptide antigen via MHC class II |
| GO:0032613 | 2.665e-02 | 0.03289 | 2
FCER1G, SASH3 | 4 | interleukin-10 production |
| GO:0032653 | 2.665e-02 | 0.03289 | 2
FCER1G, SASH3 | 4 | regulation of interleukin-10 production |
| GO:0006909 | 3.701e-02 | 0.1727 | 3
CLEC7A, FCER1G, MERTK | 21 | phagocytosis |
| GO:0019884 | 3.780e-02 | 0.04112 | 2
CTSE, FCER1G | 5 | antigen processing and presentation of exogenous antigen |
| GO:0045921 | 3.780e-02 | 0.04112 | 2
FCER1G, SEPT5 | 5 | positive regulation of exocytosis |
| GO:0002702 | 4.895e-02 | 0.04934 | 2
FCER1G, SASH3 | 6 | positive regulation of production of molecular mediator of immune response |
| GO:0002720 | 4.895e-02 | 0.04934 | 2
FCER1G, SASH3 | 6 | positive regulation of cytokine production during immune response |
Help |
Hide |
Top
Help |
Show |
Top
GO CC test for over-representation
HELP
List of all enriched GO categories (cellular components), at the 0.05
p-value level.
The columns:
- ExpCount is the expected count of genes in the
module annotated with the given GO term, just by chance.
- Count
is the number of genes in the module annotated with the given GO
term.
- Size is the total number of genes (in our universe)
annotated with the GO term.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
GO category.
Clicking on the GO identifiers takes you to the Gene Ontology web
pages.
— Click on the Help button again to close this help window.
| Id | Pvalue | ExpCount | Count | Size | Term |
| GO:0005576 | 2.817e-02 | 6.269 | 15
BCAN, C2orf72, COL11A2, DSPP, F11, FGL1, IGHG1, IL16, MATN3, PRB3, RLN1, SAA4, SPARCL1, WFDC2, WNT2B | 700 | extracellular region |
| GO:0044421 | 4.819e-02 | 3.466 | 10
BCAN, COL11A2, DSPP, F11, FGL1, IL16, MATN3, SPARCL1, WFDC2, WNT2B | 387 | extracellular region part |
Help |
Hide |
Top
Help |
Show |
Top
GO MF test for over-representation
HELP
List of all enriched GO categories (molecular function), at the 0.05
p-value level.
The columns:
- ExpCount is the expected count of genes in the
module annotated with the given GO term, just by chance.
- Count
is the number of genes in the module annotated with the given GO
term.
- Size is the total number of genes (in our universe)
annotated with the GO term.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
GO category.
Clicking on the GO identifiers takes you to the Gene Ontology web
pages.
— Click on the Help button again to close this help window.
Help |
Hide |
Top
Help |
Show |
Top
KEGG Pathway test for over-representation
HELP
List of all enriched KEGG pathways, at the 0.05
p-value level.
The columns:
- ExpCount is the expected count of genes in the
module annotated with the given KEGG pathway, just by chance.
- Count
is the number of genes in the module annotated with the given KEGG
pathway.
- Size is the total number of genes (in our universe)
annotated with the KEGG pathway.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
KEGG pathway.
Clicking on the KEGG identifiers takes you to the KEGG web site.
— Click on the Help button again to close this help window.
Help |
Hide |
Top
Help |
Show |
Top
miRNA test for over-representation
HELP
List of all enriched miRNA families, at the 0.05
p-value level.
The columns:
- ExpCount is the expected count of genes in the
module regulated by the given miRNA family, just by chance.
- Count
is the number of genes in the module regulated by the given miRNA
family.
- Size is the total number of genes (in our universe)
regulated with the given miRNA family.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
miRNA family.
The miRNA regulation data was taken from the TargetScan database.
(Only the conserved sites were used for the current analysis.)
Clicking on the miRNA names takes you to the TargetScan web site.
— Click on the Help button again to close this help window.
Help |
Hide |
Top
Help |
Show |
Top
Chromosome test for over-representation
HELP
List of all enriched Chromosomes, at the 0.05
p-value level.
The columns:
- ExpCount is the expected number of genes in the
module on the given chromosome, just by chance.
- Count
is the number of genes in the module on the given chromosome.
- Size is the total number of genes (in our universe)
on the given chromosome.
Clicking on Count shows the genes that drive the
enrichment. You can also click on the individual numbers in
the Count column, to show the driving genes for that individual
chromosome.
— Click on the Help button again to close this help window.
HELP
A list of all genes in the current module, in alphabetical order. The
size of the text corresponds to the gene scores.
Note that some gene symbols may show up more than once, if many
probes match the same Entrez gene.
Genes with no Entrez mapping are given separately, with their
Affymetrics probe ID.
— Click on the Help button again to close this help window.
Entrez genes
ACRV1acrosomal vesicle protein 1 (207969_x_at), score: 0.72
AFF2AF4/FMR2 family, member 2 (206105_at), score: 0.71
ALDH1L1aldehyde dehydrogenase 1 family, member L1 (215798_at), score: 0.7
ANK1ankyrin 1, erythrocytic (208353_x_at), score: 0.76
ARHGEF15Rho guanine nucleotide exchange factor (GEF) 15 (217348_x_at), score: 0.77
ATHL1ATH1, acid trehalase-like 1 (yeast) (219359_at), score: 0.72
ATP2C2ATPase, Ca++ transporting, type 2C, member 2 (214798_at), score: 0.67
BCANbrevican (91920_at), score: 0.75
BTNL3butyrophilin-like 3 (217207_s_at), score: 0.73
C19orf40chromosome 19 open reading frame 40 (214816_x_at), score: 0.66
C1orf61chromosome 1 open reading frame 61 (205103_at), score: 1
C2orf72chromosome 2 open reading frame 72 (213143_at), score: 0.64
C8orf60chromosome 8 open reading frame 60 (220712_at), score: 0.7
CD22CD22 molecule (38521_at), score: 0.7
CLEC7AC-type lectin domain family 7, member A (221698_s_at), score: 0.76
COL11A2collagen, type XI, alpha 2 (216993_s_at), score: 0.67
CTSEcathepsin E (205927_s_at), score: 0.64
CYTH4cytohesin 4 (219183_s_at), score: 0.65
DESdesmin (202222_s_at), score: 0.92
DSPPdentin sialophosphoprotein (221681_s_at), score: 0.84
EDARectodysplasin A receptor (220048_at), score: 0.67
ELA3Aelastase 3A, pancreatic (211738_x_at), score: 0.73
EPB41L5erythrocyte membrane protein band 4.1 like 5 (220977_x_at), score: 0.75
F11coagulation factor XI (206610_s_at), score: 0.71
FCER1GFc fragment of IgE, high affinity I, receptor for; gamma polypeptide (204232_at), score: 0.86
FGL1fibrinogen-like 1 (205305_at), score: 0.66
FMO4flavin containing monooxygenase 4 (206263_at), score: 0.66
FOLH1folate hydrolase (prostate-specific membrane antigen) 1 (217487_x_at), score: 0.7
GFERgrowth factor, augmenter of liver regeneration (204659_s_at), score: -0.65
GP1BAglycoprotein Ib (platelet), alpha polypeptide (207389_at), score: 0.64
GPR144G protein-coupled receptor 144 (216289_at), score: 0.81
HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: 0.71
HBBhemoglobin, beta (211696_x_at), score: 0.82
ICAM2intercellular adhesion molecule 2 (204683_at), score: 0.67
IGHG1immunoglobulin heavy constant gamma 1 (G1m marker) (217039_x_at), score: 0.64
IL16interleukin 16 (lymphocyte chemoattractant factor) (209827_s_at), score: 0.66
KSR1kinase suppressor of ras 1 (213769_at), score: 0.64
LOC149501similar to keratin 8 (216821_at), score: 0.64
LOC729806similar to hCG1725380 (217544_at), score: 0.71
LOC80054hypothetical LOC80054 (220465_at), score: 0.64
MALmal, T-cell differentiation protein (204777_s_at), score: 0.69
MAP3K7IP1mitogen-activated protein kinase kinase kinase 7 interacting protein 1 (203901_at), score: 0.76
MATN3matrilin 3 (206091_at), score: -0.78
MERTKc-mer proto-oncogene tyrosine kinase (211913_s_at), score: 0.65
MICAL3microtubule associated monoxygenase, calponin and LIM domain containing 3 (212715_s_at), score: -0.73
MNDAmyeloid cell nuclear differentiation antigen (204959_at), score: 0.64
MYO5Cmyosin VC (218966_at), score: 0.64
NCAM2neural cell adhesion molecule 2 (205669_at), score: 0.77
NCR2natural cytotoxicity triggering receptor 2 (217045_x_at), score: 0.65
NOVA2neuro-oncological ventral antigen 2 (206477_s_at), score: 0.67
OCLMoculomedin (208274_at), score: 0.79
OR7E47Polfactory receptor, family 7, subfamily E, member 47 pseudogene (216698_x_at), score: 0.65
PMS2L2postmeiotic segregation increased 2-like 2 pseudogene (215410_at), score: 0.72
PRB3proline-rich protein BstNI subfamily 3 (206998_x_at), score: 0.65
PTPN6protein tyrosine phosphatase, non-receptor type 6 (206687_s_at), score: 0.67
PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: 0.74
RLN1relaxin 1 (211753_s_at), score: 0.69
ROS1c-ros oncogene 1 , receptor tyrosine kinase (207569_at), score: 0.69
RP3-377H14.5hypothetical LOC285830 (222279_at), score: 0.64
RPS17P5ribosomal protein S17 pseudogene 5 (216348_at), score: 0.65
RRP8ribosomal RNA processing 8, methyltransferase, homolog (yeast) (203171_s_at), score: -0.68
RUFY2RUN and FYVE domain containing 2 (219957_at), score: 0.8
SAA4serum amyloid A4, constitutive (207096_at), score: 0.73
SASH3SAM and SH3 domain containing 3 (204923_at), score: 0.68
SEPT5septin 5 (209767_s_at), score: 0.75
SERPINB3serpin peptidase inhibitor, clade B (ovalbumin), member 3 (209719_x_at), score: 0.65
SH2D3ASH2 domain containing 3A (222169_x_at), score: 0.72
SLC1A7solute carrier family 1 (glutamate transporter), member 7 (210923_at), score: 0.82
SPARCL1SPARC-like 1 (hevin) (200795_at), score: 0.67
TAF4BTAF4b RNA polymerase II, TATA box binding protein (TBP)-associated factor, 105kDa (216226_at), score: 0.73
TAS2R14taste receptor, type 2, member 14 (221391_at), score: 0.89
TLE6transducin-like enhancer of split 6 (E(sp1) homolog, Drosophila) (222219_s_at), score: 0.71
TMC5transmembrane channel-like 5 (219580_s_at), score: 0.84
TMEM144transmembrane protein 144 (219750_at), score: 0.74
TMSB4Ythymosin beta 4, Y-linked (206769_at), score: 0.69
TNFRSF8tumor necrosis factor receptor superfamily, member 8 (206729_at), score: 0.7
VHLvon Hippel-Lindau tumor suppressor (203844_at), score: 0.66
WFDC2WAP four-disulfide core domain 2 (203892_at), score: 0.92
WNT2Bwingless-type MMTV integration site family, member 2B (206459_s_at), score: 0.86
ZDHHC11zinc finger, DHHC-type containing 11 (221646_s_at), score: 0.7
ZNF37Azinc finger protein 37A (214878_at), score: 0.75
Non-Entrez genes
Unknown, score:
HELP
Conditions in the module, given in the same order as on the expression
plot above. Red color means over-expression, green under-expression in
the given condition.
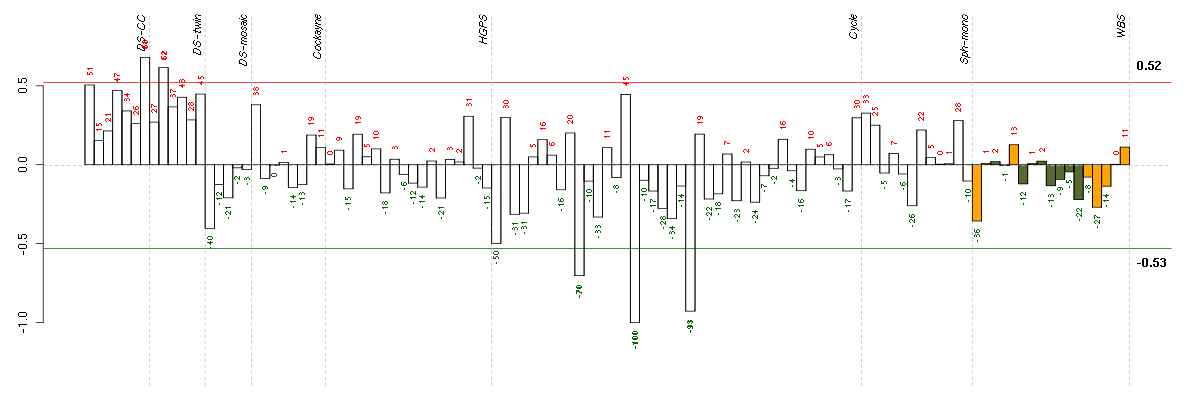
The barplot below shows the condition (sample) scores. A separate bar
is shown for each sample, its height is the corresponding score of the
sample in the module. The red and green numbers on the bars are the
sample scores expressed in percents, i.e. 100% is 1.0.
The red and green lines show the module thresholds, samples above
the red line and below the green line are included in the module.
The different experiments that were part of the study, are separated
by dashed vertical lines.
— Click on the Help button again to close this help window.
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
| E-TABM-263-raw-cel-1515485951.cel | 16 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |



© 2008-2010 Computational Biology Group, Department of Medical Genetics,
University of Lausanne, Switzerland