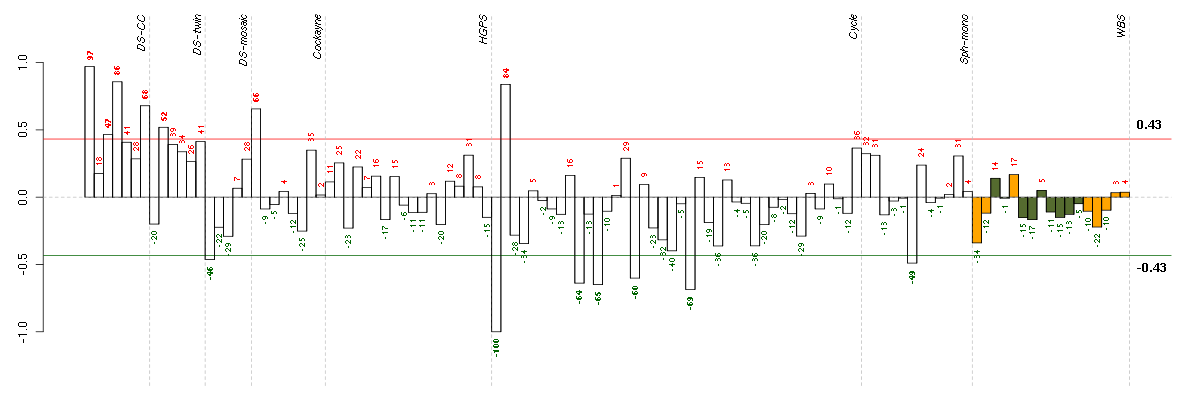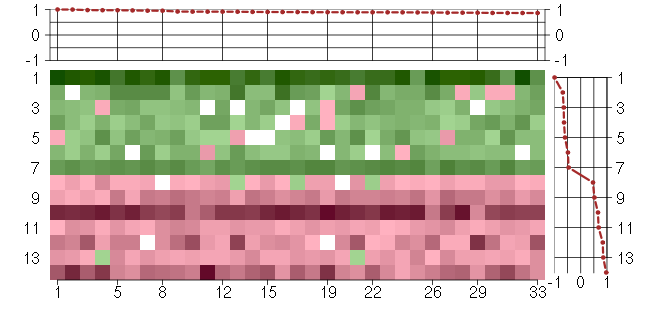

Under-expression is coded with green,
over-expression with red color.


extracellular region
The space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers the host cell environment outside an intracellular parasite.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.
all
This term is the most general term possible
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.

A1CFAPOBEC1 complementation factor (220951_s_at), score: 0.89 ADAMTS7ADAM metallopeptidase with thrombospondin type 1 motif, 7 (220705_s_at), score: 0.92 BMP10bone morphogenetic protein 10 (208292_at), score: 0.9 BTNL3butyrophilin-like 3 (217207_s_at), score: 0.95 C21orf2chromosome 21 open reading frame 2 (203996_s_at), score: 0.91 C7orf28Achromosome 7 open reading frame 28A (201974_s_at), score: 0.86 CCDC9coiled-coil domain containing 9 (206257_at), score: 0.87 CLDN10claudin 10 (205328_at), score: 0.9 COL11A2collagen, type XI, alpha 2 (216993_s_at), score: 0.97 DHRS9dehydrogenase/reductase (SDR family) member 9 (219799_s_at), score: 0.87 DMBT1deleted in malignant brain tumors 1 (208250_s_at), score: 0.89 DNASE1L3deoxyribonuclease I-like 3 (205554_s_at), score: 0.89 HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: 0.89 HSPA6heat shock 70kDa protein 6 (HSP70B') (117_at), score: 0.86 KRT8P12keratin 8 pseudogene 12 (222060_at), score: 0.87 LOC100188945cell division cycle associated 4 pseudogene (215109_at), score: 0.91 LOC80054hypothetical LOC80054 (220465_at), score: 0.88 MMRN2multimerin 2 (219091_s_at), score: 0.89 MOBPmyelin-associated oligodendrocyte basic protein (210193_at), score: 1 MYL4myosin, light chain 4, alkali; atrial, embryonic (210088_x_at), score: 0.86 MYO1Amyosin IA (211916_s_at), score: 0.97 PECAM1platelet/endothelial cell adhesion molecule (208982_at), score: 0.88 PGK2phosphoglycerate kinase 2 (217009_at), score: 0.98 PON1paraoxonase 1 (206344_at), score: 0.88 PRSS7protease, serine, 7 (enterokinase) (217269_s_at), score: 1 PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: 0.95 ROS1c-ros oncogene 1 , receptor tyrosine kinase (207569_at), score: 0.92 RPGRIP1retinitis pigmentosa GTPase regulator interacting protein 1 (206608_s_at), score: 0.91 S100A14S100 calcium binding protein A14 (218677_at), score: 0.9 SLC26A10solute carrier family 26, member 10 (214951_at), score: 0.89 TBX21T-box 21 (220684_at), score: 0.9 TMSB4Ythymosin beta 4, Y-linked (206769_at), score: 0.96 VGFVGF nerve growth factor inducible (205586_x_at), score: 0.91
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485951.cel | 16 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-4219-raw-cel-1311956358.cel | 10 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| ctrl c 08-03.CEL | 3 | 1 | DS-CC | hgu133a | none | DS-CC 3 |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| E-TABM-263-raw-cel-1515485671.cel | 2 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |