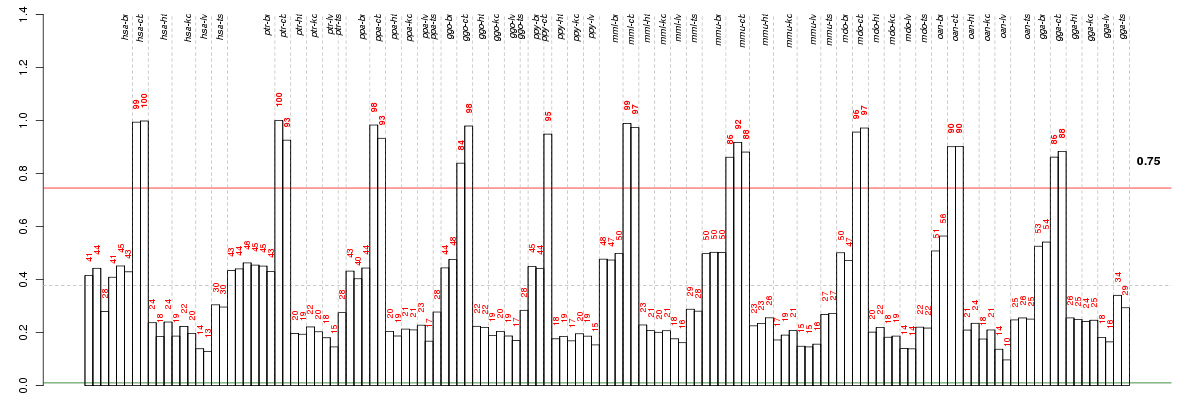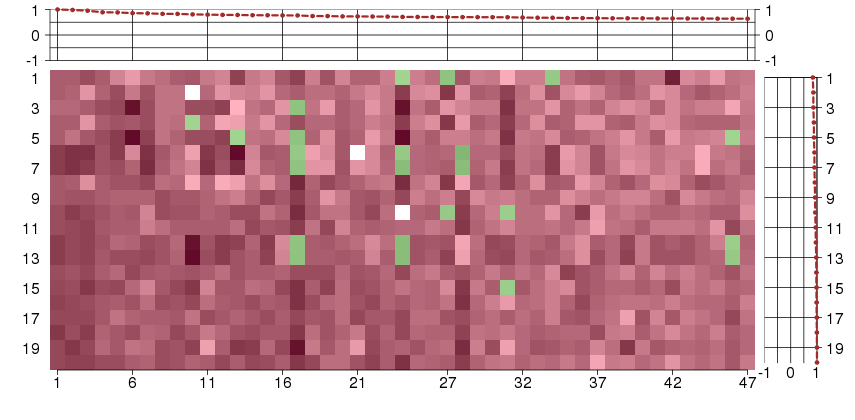

Under-expression is coded with green,
over-expression with red color.



C15orf27chromosome 15 open reading frame 27 (ENSG00000169758), score: 0.7 C7orf16chromosome 7 open reading frame 16 (ENSG00000106341), score: 0.71 C8orf79chromosome 8 open reading frame 79 (ENSG00000170941), score: 0.74 CA8carbonic anhydrase VIII (ENSG00000178538), score: 0.7 CBLN1cerebellin 1 precursor (ENSG00000102924), score: 0.89 CDH7cadherin 7, type 2 (ENSG00000081138), score: 0.74 CHN2chimerin (chimaerin) 2 (ENSG00000106069), score: 0.65 CRTAMcytotoxic and regulatory T cell molecule (ENSG00000109943), score: 0.77 EPC1enhancer of polycomb homolog 1 (Drosophila) (ENSG00000120616), score: 0.65 FAT2FAT tumor suppressor homolog 2 (Drosophila) (ENSG00000086570), score: 1 FGF5fibroblast growth factor 5 (ENSG00000138675), score: 0.81 GALNT7UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 7 (GalNAc-T7) (ENSG00000109586), score: 0.67 GNG13guanine nucleotide binding protein (G protein), gamma 13 (ENSG00000127588), score: 0.68 GRIA4glutamate receptor, ionotrophic, AMPA 4 (ENSG00000152578), score: 0.65 GRID2glutamate receptor, ionotropic, delta 2 (ENSG00000152208), score: 0.85 GRM1glutamate receptor, metabotropic 1 (ENSG00000152822), score: 0.73 KCNC1potassium voltage-gated channel, Shaw-related subfamily, member 1 (ENSG00000129159), score: 0.68 KCNK10potassium channel, subfamily K, member 10 (ENSG00000100433), score: 0.73 KCNK9potassium channel, subfamily K, member 9 (ENSG00000169427), score: 0.77 LRCH1leucine-rich repeats and calponin homology (CH) domain containing 1 (ENSG00000136141), score: 0.66 LRRC38leucine rich repeat containing 38 (ENSG00000162494), score: 0.66 MAB21L1mab-21-like 1 (C. elegans) (ENSG00000180660), score: 0.98 MAML3mastermind-like 3 (Drosophila) (ENSG00000196782), score: 0.79 MDGA1MAM domain containing glycosylphosphatidylinositol anchor 1 (ENSG00000112139), score: 0.7 MEIS1Meis homeobox 1 (ENSG00000143995), score: 0.65 MYT1myelin transcription factor 1 (ENSG00000196132), score: 0.78 NRG2neuregulin 2 (ENSG00000158458), score: 0.7 PAX3paired box 3 (ENSG00000135903), score: 0.65 PAX6paired box 6 (ENSG00000007372), score: 0.83 PLCB4phospholipase C, beta 4 (ENSG00000101333), score: 0.7 PLD5phospholipase D family, member 5 (ENSG00000180287), score: 0.64 PTCH1patched 1 (ENSG00000185920), score: 0.64 PTCHD1patched domain containing 1 (ENSG00000165186), score: 0.72 PVALBparvalbumin (ENSG00000100362), score: 0.64 SHFSrc homology 2 domain containing F (ENSG00000138606), score: 0.65 SKOR1SKI family transcriptional corepressor 1 (ENSG00000188779), score: 0.77 SLC35F4solute carrier family 35, member F4 (ENSG00000151812), score: 0.79 SPHKAPSPHK1 interactor, AKAP domain containing (ENSG00000153820), score: 0.65 TIAM1T-cell lymphoma invasion and metastasis 1 (ENSG00000156299), score: 0.72 TLL1tolloid-like 1 (ENSG00000038295), score: 0.86 TRIM67tripartite motif-containing 67 (ENSG00000119283), score: 0.89 TRPC3transient receptor potential cation channel, subfamily C, member 3 (ENSG00000138741), score: 0.7 UPF0639UPF0639 protein (ENSG00000175985), score: 0.8 ZFPM2zinc finger protein, multitype 2 (ENSG00000169946), score: 0.68 ZIC4Zic family member 4 (ENSG00000174963), score: 0.95 ZNF238zinc finger protein 238 (ENSG00000179456), score: 0.71 ZNF521zinc finger protein 521 (ENSG00000198795), score: 0.83
| Id | species | tissue | sex | individual |
|---|---|---|---|---|
| ggo_cb_m_ca1 | ggo | cb | m | _ |
| mmu_cb_m1_ca1 | mmu | cb | m | 1 |
| gga_cb_m_ca1 | gga | cb | m | _ |
| mmu_cb_f_ca1 | mmu | cb | f | _ |
| gga_cb_f_ca1 | gga | cb | f | _ |
| oan_cb_m_ca1 | oan | cb | m | _ |
| oan_cb_f_ca1 | oan | cb | f | _ |
| mmu_cb_m2_ca1 | mmu | cb | m | 2 |
| ptr_cb_f_ca1 | ptr | cb | f | _ |
| ppa_cb_f_ca1 | ppa | cb | f | _ |
| ppy_cb_f_ca1 | ppy | cb | f | _ |
| mdo_cb_m_ca1 | mdo | cb | m | _ |
| mdo_cb_f_ca1 | mdo | cb | f | _ |
| mml_cb_f_ca1 | mml | cb | f | _ |
| ggo_cb_f_ca1 | ggo | cb | f | _ |
| ppa_cb_m_ca1 | ppa | cb | m | _ |
| mml_cb_m_ca1 | mml | cb | m | _ |
| hsa_cb_m_ca1 | hsa | cb | m | _ |
| hsa_cb_f_ca1 | hsa | cb | f | _ |
| ptr_cb_m_ca1 | ptr | cb | m | _ |