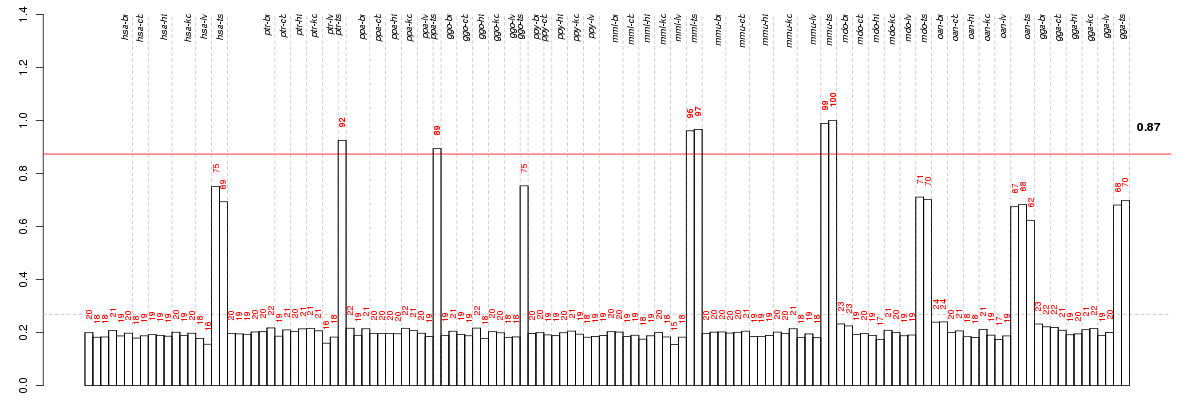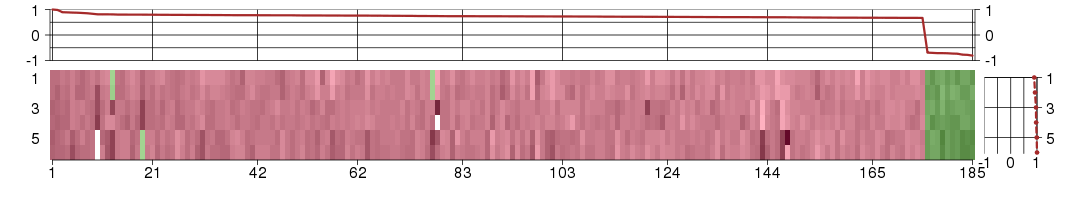

Under-expression is coded with green,
over-expression with red color.

reproduction
The production by an organism of new individuals that contain some portion of their genetic material inherited from that organism.
M phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the mitotic cell cycle during which mitosis takes place.
mitotic cell cycle
Progression through the phases of the mitotic cell cycle, the most common eukaryotic cell cycle, which canonically comprises four successive phases called G1, S, G2, and M and includes replication of the genome and the subsequent segregation of chromosomes into daughter cells. In some variant cell cycles nuclear replication or nuclear division may not be followed by cell division, or G1 and G2 phases may be absent.
M phase
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the cell cycle comprising nuclear division.
nuclear division
A process by which a cell nucleus is divided into two nuclei, with DNA and other nuclear contents distributed between the daughter nuclei.
organelle organization
A process that is carried out at the cellular level which results in the assembly, arrangement of constituent parts, or disassembly of an organelle within a cell. An organelle is an organized structure of distinctive morphology and function. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
microtubule-based process
Any cellular process that depends upon or alters the microtubule cytoskeleton, that part of the cytoskeleton comprising microtubules and their associated proteins.
cell cycle
The progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events. Canonically, the cell cycle comprises the replication and segregation of genetic material followed by the division of the cell, but in endocycles or syncytial cells nuclear replication or nuclear division may not be followed by cell division.
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
meiosis
A cell cycle process comprising the steps by which a cell progresses through the nuclear division phase of a meiotic cell cycle, the specialized nuclear and cell division in which a single diploid cell undergoes two nuclear divisions following a single round of DNA replication in order to produce four daughter cells that contain half the number of chromosomes as the diploid cell. Meiotic division occurs during the formation of gametes from diploid organisms and at the beginning of haplophase in those organisms that alternate between diploid and haploid generations.
gamete generation
The generation and maintenance of gametes in a multicellular organism. A gamete is a haploid reproductive cell.
spermatogenesis
The process of formation of spermatozoa, including spermatocytogenesis and spermiogenesis.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
fertilization
The union of gametes of opposite sexes during the process of sexual reproduction to form a zygote. It involves the fusion of the gametic nuclei (karyogamy) and cytoplasm (plasmogamy).
cellular process
Any process that is carried out at the cellular level, but not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
cellular component organization
A process that is carried out at the cellular level which results in the assembly, arrangement of constituent parts, or disassembly of a cellular component.
sexual reproduction
The regular alternation, in the life cycle of haplontic, diplontic and diplohaplontic organisms, of meiosis and fertilization which provides for the production offspring. In diplontic organisms there is a life cycle in which the products of meiosis behave directly as gametes, fusing to form a zygote from which the diploid, or sexually reproductive polyploid, adult organism will develop. In diplohaplontic organisms a haploid phase (gametophyte) exists in the life cycle between meiosis and fertilization (e.g. higher plants, many algae and Fungi); the products of meiosis are spores that develop as haploid individuals from which haploid gametes develop to form a diploid zygote; diplohaplontic organisms show an alternation of haploid and diploid generations. In haplontic organisms meiosis occurs in the zygote, giving rise to four haploid cells (e.g. many algae and protozoa), only the zygote is diploid and this may form a resistant spore, tiding organisms over hard times.
cell cycle process
A cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
cell cycle phase
A cell cycle process comprising the steps by which a cell progresses through one of the biochemical and morphological phases and events that occur during successive cell replication or nuclear replication events.
reproductive process
A biological process that directly contributes to the process of producing new individuals by one or two organisms. The new individuals inherit some proportion of their genetic material from the parent or parents.
multicellular organismal process
Any biological process, occurring at the level of a multicellular organism, pertinent to its function.
multicellular organism reproduction
The biological process by which new individuals are produced by one or two multicellular organisms. The new individuals inherit some proportion of their genetic material from the parent or parents.
male gamete generation
Generation of the male gamete; specialised haploid cells produced by meiosis and along with a female gamete takes part in sexual reproduction.
organelle fission
The creation of two or more organelles by division of one organelle.
reproductive process in a multicellular organism
The process, occurring above the cellular level, that is pertinent to the reproductive function of a multicellular organism. This includes the integrated processes at the level of tissues and organs.
cell division
The process resulting in the physical partitioning and separation of a cell into daughter cells.
meiotic cell cycle
Progression through the phases of the meiotic cell cycle, in which canonically a cell replicates to produce four offspring with half the chromosomal content of the progenitor cell.
M phase of meiotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the meiotic cell cycle during which meiosis takes place.
all
NA
reproductive process
A biological process that directly contributes to the process of producing new individuals by one or two organisms. The new individuals inherit some proportion of their genetic material from the parent or parents.
organelle organization
A process that is carried out at the cellular level which results in the assembly, arrangement of constituent parts, or disassembly of an organelle within a cell. An organelle is an organized structure of distinctive morphology and function. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
multicellular organism reproduction
The biological process by which new individuals are produced by one or two multicellular organisms. The new individuals inherit some proportion of their genetic material from the parent or parents.
reproductive process in a multicellular organism
The process, occurring above the cellular level, that is pertinent to the reproductive function of a multicellular organism. This includes the integrated processes at the level of tissues and organs.
fertilization
The union of gametes of opposite sexes during the process of sexual reproduction to form a zygote. It involves the fusion of the gametic nuclei (karyogamy) and cytoplasm (plasmogamy).
reproductive process in a multicellular organism
The process, occurring above the cellular level, that is pertinent to the reproductive function of a multicellular organism. This includes the integrated processes at the level of tissues and organs.
cell cycle process
A cellular process that is involved in the progression of biochemical and morphological phases and events that occur in a cell during successive cell replication or nuclear replication events.
gamete generation
The generation and maintenance of gametes in a multicellular organism. A gamete is a haploid reproductive cell.
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
mitosis
A cell cycle process comprising the steps by which the nucleus of a eukaryotic cell divides; the process involves condensation of chromosomal DNA into a highly compacted form. Canonically, mitosis produces two daughter nuclei whose chromosome complement is identical to that of the mother cell.
meiosis
A cell cycle process comprising the steps by which a cell progresses through the nuclear division phase of a meiotic cell cycle, the specialized nuclear and cell division in which a single diploid cell undergoes two nuclear divisions following a single round of DNA replication in order to produce four daughter cells that contain half the number of chromosomes as the diploid cell. Meiotic division occurs during the formation of gametes from diploid organisms and at the beginning of haplophase in those organisms that alternate between diploid and haploid generations.
M phase of mitotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the mitotic cell cycle during which mitosis takes place.
M phase of meiotic cell cycle
A cell cycle process comprising the steps by which a cell progresses through M phase, the part of the meiotic cell cycle during which meiosis takes place.

condensed chromosome
A highly compacted molecule of DNA and associated proteins resulting in a cytologically distinct structure.
intracellular
The living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
chromosome, centromeric region
The region of a chromosome that includes the centromeric DNA and associated proteins. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
condensed chromosome kinetochore
A multisubunit complex that is located at the centromeric region of a condensed chromosome and provides an attachment point for the spindle microtubules.
condensed chromosome, centromeric region
The region of a condensed chromosome that includes the centromere and associated proteins, including the kinetochore. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
spindle pole
Either of the ends of a spindle, where spindle microtubules are organized; usually contains a microtubule organizing center and accessory molecules, spindle microtubules and astral microtubules.
ribonucleoprotein complex
A macromolecular complex containing both protein and RNA molecules.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
cell
The basic structural and functional unit of all organisms. Includes the plasma membrane and any external encapsulating structures such as the cell wall and cell envelope.
cytoplasm
All of the contents of a cell excluding the plasma membrane and nucleus, but including other subcellular structures.
chromosome
A structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
microtubule organizing center
A cytoplasmic structure that can catalyze gamma-tubulin-dependent microtubule nucleation and that can anchor microtubules by interacting with their minus ends, plus ends or sides.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
cytoskeleton
Any of the various filamentous elements that form the internal framework of cells, and typically remain after treatment of the cells with mild detergent to remove membrane constituents and soluble components of the cytoplasm. The term embraces intermediate filaments, microfilaments, microtubules, the microtrabecular lattice, and other structures characterized by a polymeric filamentous nature and long-range order within the cell. The various elements of the cytoskeleton not only serve in the maintenance of cellular shape but also have roles in other cellular functions, including cellular movement, cell division, endocytosis, and movement of organelles.
microtubule
Any of the long, generally straight, hollow tubes of internal diameter 12-15 nm and external diameter 24 nm found in a wide variety of eukaryotic cells; each consists (usually) of 13 protofilaments of polymeric tubulin, staggered in such a manner that the tubulin monomers are arranged in a helical pattern on the microtubular surface, and with the alpha/beta axes of the tubulin subunits parallel to the long axis of the tubule; exist in equilibrium with pool of tubulin monomers and can be rapidly assembled or disassembled in response to physiological stimuli; concerned with force generation, e.g. in the spindle.
spindle microtubule
Any microtubule that is part of a mitotic or meiotic spindle; anchored at one spindle pole.
microtubule cytoskeleton
The part of the cytoskeleton (the internal framework of a cell) composed of microtubules and associated proteins.
macromolecular complex
A stable assembly of two or more macromolecules, i.e. proteins, nucleic acids, carbohydrates or lipids, in which the constituent parts function together.
chromatoid body
A ribonucleoprotein complex found in the cytoplasm of male germ cells, composed of exceedingly thin filaments that are consolidated into a compact mass or into dense strands of varying thickness that branch to form an irregular network. Contains mRNAs, miRNAs, and protein components involved in miRNA processing (such as Argonaute proteins and the endonuclease Dicer) and in RNA decay (such as the decapping enzyme DCP1a and GW182).
P granule
A small cytoplasmic, non-membranous RNA/protein complex aggregates in the primordial germ cells of many higher eukaryotes.
organelle
Organized structure of distinctive morphology and function. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton, and prokaryotic structures such as anammoxosomes and pirellulosomes. Excludes the plasma membrane.
non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane. Includes ribosomes, the cytoskeleton and chromosomes.
intracellular organelle
Organized structure of distinctive morphology and function, occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
intracellular non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane and occurring within the cell. Includes ribosomes, the cytoskeleton and chromosomes.
protein complex
Any macromolecular complex composed of two or more polypeptide subunits, which may or may not be identical. Protein complexes may have other associated non-protein prosthetic groups, such as nucleotides, metal ions or other small molecules.
organelle part
Any constituent part of an organelle, an organized structure of distinctive morphology and function. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton, but excludes the plasma membrane.
intracellular part
Any constituent part of the living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
chromosomal part
Any constituent part of a chromosome, a structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
cytoskeletal part
Any constituent part of the cytoskeleton, a cellular scaffolding or skeleton that maintains cell shape, enables some cell motion (using structures such as flagella and cilia), and plays important roles in both intra-cellular transport (e.g. the movement of vesicles and organelles) and cellular division. Includes constituent parts of intermediate filaments, microfilaments, microtubules, and the microtrabecular lattice.
cytoplasmic part
Any constituent part of the cytoplasm, all of the contents of a cell excluding the plasma membrane and nucleus, but including other subcellular structures.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
pole plasm
Differentiated cytoplasm associated with a pole (animal, vegetal, anterior, or posterior) of an oocyte, egg or early embryo.
germ plasm
Differentiated cytoplasm associated with a pole of an oocyte, egg or early embryo that will be inherited by the cells that will give rise to the germ line.
all
NA
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
organelle part
Any constituent part of an organelle, an organized structure of distinctive morphology and function. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton, but excludes the plasma membrane.
intracellular non-membrane-bounded organelle
Organized structure of distinctive morphology and function, not bounded by a lipid bilayer membrane and occurring within the cell. Includes ribosomes, the cytoskeleton and chromosomes.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
intracellular part
Any constituent part of the living contents of a cell; the matter contained within (but not including) the plasma membrane, usually taken to exclude large vacuoles and masses of secretory or ingested material. In eukaryotes it includes the nucleus and cytoplasm.
ribonucleoprotein complex
A macromolecular complex containing both protein and RNA molecules.
intracellular organelle
Organized structure of distinctive morphology and function, occurring within the cell. Includes the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton. Excludes the plasma membrane.
intracellular organelle part
A constituent part of an intracellular organelle, an organized structure of distinctive morphology and function, occurring within the cell. Includes constituent parts of the nucleus, mitochondria, plastids, vacuoles, vesicles, ribosomes and the cytoskeleton but excludes the plasma membrane.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
microtubule
Any of the long, generally straight, hollow tubes of internal diameter 12-15 nm and external diameter 24 nm found in a wide variety of eukaryotic cells; each consists (usually) of 13 protofilaments of polymeric tubulin, staggered in such a manner that the tubulin monomers are arranged in a helical pattern on the microtubular surface, and with the alpha/beta axes of the tubulin subunits parallel to the long axis of the tubule; exist in equilibrium with pool of tubulin monomers and can be rapidly assembled or disassembled in response to physiological stimuli; concerned with force generation, e.g. in the spindle.
cytoplasmic part
Any constituent part of the cytoplasm, all of the contents of a cell excluding the plasma membrane and nucleus, but including other subcellular structures.
microtubule organizing center
A cytoplasmic structure that can catalyze gamma-tubulin-dependent microtubule nucleation and that can anchor microtubules by interacting with their minus ends, plus ends or sides.
P granule
A small cytoplasmic, non-membranous RNA/protein complex aggregates in the primordial germ cells of many higher eukaryotes.
chromosomal part
Any constituent part of a chromosome, a structure composed of a very long molecule of DNA and associated proteins (e.g. histones) that carries hereditary information.
spindle pole
Either of the ends of a spindle, where spindle microtubules are organized; usually contains a microtubule organizing center and accessory molecules, spindle microtubules and astral microtubules.
spindle microtubule
Any microtubule that is part of a mitotic or meiotic spindle; anchored at one spindle pole.
cytoskeletal part
Any constituent part of the cytoskeleton, a cellular scaffolding or skeleton that maintains cell shape, enables some cell motion (using structures such as flagella and cilia), and plays important roles in both intra-cellular transport (e.g. the movement of vesicles and organelles) and cellular division. Includes constituent parts of intermediate filaments, microfilaments, microtubules, and the microtrabecular lattice.
kinetochore
A multisubunit complex that is located at the centromeric region of DNA and provides an attachment point for the spindle microtubules.
condensed chromosome, centromeric region
The region of a condensed chromosome that includes the centromere and associated proteins, including the kinetochore. In monocentric chromosomes, this region corresponds to a single area of the chromosome, whereas in holocentric chromosomes, it is evenly distributed along the chromosome.
microtubule organizing center
A cytoplasmic structure that can catalyze gamma-tubulin-dependent microtubule nucleation and that can anchor microtubules by interacting with their minus ends, plus ends or sides.
spindle
The array of microtubules and associated molecules that forms between opposite poles of a eukaryotic cell during mitosis or meiosis and serves to move the duplicated chromosomes apart.
microtubule
Any of the long, generally straight, hollow tubes of internal diameter 12-15 nm and external diameter 24 nm found in a wide variety of eukaryotic cells; each consists (usually) of 13 protofilaments of polymeric tubulin, staggered in such a manner that the tubulin monomers are arranged in a helical pattern on the microtubular surface, and with the alpha/beta axes of the tubulin subunits parallel to the long axis of the tubule; exist in equilibrium with pool of tubulin monomers and can be rapidly assembled or disassembled in response to physiological stimuli; concerned with force generation, e.g. in the spindle.
condensed chromosome kinetochore
A multisubunit complex that is located at the centromeric region of a condensed chromosome and provides an attachment point for the spindle microtubules.
P granule
A small cytoplasmic, non-membranous RNA/protein complex aggregates in the primordial germ cells of many higher eukaryotes.

molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
motor activity
Catalysis of movement along a polymeric molecule such as a microfilament or microtubule, coupled to the hydrolysis of a nucleoside triphosphate.
microtubule motor activity
Catalysis of movement along a microtubule, coupled to the hydrolysis of a nucleoside triphosphate (usually ATP).
catalytic activity
Catalysis of a biochemical reaction at physiological temperatures. In biologically catalyzed reactions, the reactants are known as substrates, and the catalysts are naturally occurring macromolecular substances known as enzymes. Enzymes possess specific binding sites for substrates, and are usually composed wholly or largely of protein, but RNA that has catalytic activity (ribozyme) is often also regarded as enzymatic.
nucleoside-triphosphatase activity
Catalysis of the reaction: a nucleoside triphosphate + H2O = nucleoside diphosphate + phosphate.
pyrophosphatase activity
Catalysis of the hydrolysis of a pyrophosphate bond between two phosphate groups, leaving one phosphate on each of the two fragments.
hydrolase activity
Catalysis of the hydrolysis of various bonds, e.g. C-O, C-N, C-C, phosphoric anhydride bonds, etc. Hydrolase is the systematic name for any enzyme of EC class 3.
hydrolase activity, acting on acid anhydrides
Catalysis of the hydrolysis of any acid anhydride.
hydrolase activity, acting on acid anhydrides, in phosphorus-containing anhydrides
Catalysis of the hydrolysis of any acid anhydride which contains phosphorus.
all
NA
ACBD6acyl-CoA binding domain containing 6 (ENSG00000135847), score: 0.8 ADAD1adenosine deaminase domain containing 1 (testis-specific) (ENSG00000164113), score: 0.8 AGBL3ATP/GTP binding protein-like 3 (ENSG00000146856), score: 0.68 AGBL5ATP/GTP binding protein-like 5 (ENSG00000084693), score: 0.69 AHCTF1AT hook containing transcription factor 1 (ENSG00000153207), score: 0.71 AK7adenylate kinase 7 (ENSG00000140057), score: 0.78 ANKRD5ankyrin repeat domain 5 (ENSG00000132623), score: 0.77 ARID2AT rich interactive domain 2 (ARID, RFX-like) (ENSG00000189079), score: 0.73 ARMC3armadillo repeat containing 3 (ENSG00000165309), score: 0.76 ARMC4armadillo repeat containing 4 (ENSG00000169126), score: 0.75 ASPHaspartate beta-hydroxylase (ENSG00000198363), score: -0.81 AURKAaurora kinase A (ENSG00000087586), score: 0.77 BAG5BCL2-associated athanogene 5 (ENSG00000166170), score: 0.73 BBXbobby sox homolog (Drosophila) (ENSG00000114439), score: 0.71 BOLLbol, boule-like (Drosophila) (ENSG00000152430), score: 0.76 BRIP1BRCA1 interacting protein C-terminal helicase 1 (ENSG00000136492), score: 0.73 BTG4B-cell translocation gene 4 (ENSG00000137707), score: 0.77 BUB1budding uninhibited by benzimidazoles 1 homolog (yeast) (ENSG00000169679), score: 0.7 C10orf46chromosome 10 open reading frame 46 (ENSG00000151893), score: 0.7 C10orf96chromosome 10 open reading frame 96 (ENSG00000182645), score: 0.89 C11orf70chromosome 11 open reading frame 70 (ENSG00000137691), score: 0.68 C11orf82chromosome 11 open reading frame 82 (ENSG00000165490), score: 0.76 C12orf50chromosome 12 open reading frame 50 (ENSG00000165805), score: 0.68 C12orf63chromosome 12 open reading frame 63 (ENSG00000188596), score: 0.71 C14orf166Bchromosome 14 open reading frame 166B (ENSG00000100565), score: 0.79 C14orf50chromosome 14 open reading frame 50 (ENSG00000165807), score: 0.71 C15orf26chromosome 15 open reading frame 26 (ENSG00000156206), score: 0.78 C19orf45chromosome 19 open reading frame 45 (ENSG00000198723), score: 0.76 C1orf100chromosome 1 open reading frame 100 (ENSG00000173728), score: 0.71 C1orf111chromosome 1 open reading frame 111 (ENSG00000171722), score: 0.67 C1orf158chromosome 1 open reading frame 158 (ENSG00000157330), score: 0.7 C20orf85chromosome 20 open reading frame 85 (ENSG00000124237), score: 0.72 C2CD3C2 calcium-dependent domain containing 3 (ENSG00000168014), score: 0.67 C2orf71chromosome 2 open reading frame 71 (ENSG00000179270), score: 0.69 C3orf38chromosome 3 open reading frame 38 (ENSG00000179021), score: 0.76 C6orf204chromosome 6 open reading frame 204 (ENSG00000111860), score: 0.68 C7orf31chromosome 7 open reading frame 31 (ENSG00000153790), score: 0.76 C7orf45chromosome 7 open reading frame 45 (ENSG00000165120), score: 0.76 C7orf62chromosome 7 open reading frame 62 (ENSG00000164645), score: 0.77 C9orf96chromosome 9 open reading frame 96 (ENSG00000198870), score: 0.72 C9orf98chromosome 9 open reading frame 98 (ENSG00000165695), score: 0.69 CALR3calreticulin 3 (ENSG00000141979), score: 0.75 CCDC108coiled-coil domain containing 108 (ENSG00000181378), score: 0.73 CCDC122coiled-coil domain containing 122 (ENSG00000151773), score: 0.68 CCDC135coiled-coil domain containing 135 (ENSG00000159625), score: 0.67 CCDC146coiled-coil domain containing 146 (ENSG00000135205), score: 0.73 CCDC30coiled-coil domain containing 30 (ENSG00000186409), score: 0.68 CCDC63coiled-coil domain containing 63 (ENSG00000173093), score: 0.79 CCDC67coiled-coil domain containing 67 (ENSG00000165325), score: 0.69 CCDC83coiled-coil domain containing 83 (ENSG00000150676), score: 0.89 CCR4chemokine (C-C motif) receptor 4 (ENSG00000183813), score: 0.81 CDC14ACDC14 cell division cycle 14 homolog A (S. cerevisiae) (ENSG00000079335), score: 0.71 CDKN3cyclin-dependent kinase inhibitor 3 (ENSG00000100526), score: 0.73 CENPFcentromere protein F, 350/400kDa (mitosin) (ENSG00000117724), score: 0.74 CEP350centrosomal protein 350kDa (ENSG00000135837), score: 0.74 CEP55centrosomal protein 55kDa (ENSG00000138180), score: 0.7 CLCA2chloride channel accessory 2 (ENSG00000137975), score: 0.81 CNOT1CCR4-NOT transcription complex, subunit 1 (ENSG00000125107), score: 0.75 CUL3cullin 3 (ENSG00000036257), score: 0.68 DCAF5DDB1 and CUL4 associated factor 5 (ENSG00000139990), score: 0.75 DDX20DEAD (Asp-Glu-Ala-Asp) box polypeptide 20 (ENSG00000064703), score: 0.79 DDX4DEAD (Asp-Glu-Ala-Asp) box polypeptide 4 (ENSG00000152670), score: 0.74 DIS3LDIS3 mitotic control homolog (S. cerevisiae)-like (ENSG00000166938), score: 0.73 DNAH8dynein, axonemal, heavy chain 8 (ENSG00000124721), score: 0.88 DNAI2dynein, axonemal, intermediate chain 2 (ENSG00000171595), score: 0.72 DNAJC21DnaJ (Hsp40) homolog, subfamily C, member 21 (ENSG00000168724), score: 0.74 DNAJC5BDnaJ (Hsp40) homolog, subfamily C, member 5 beta (ENSG00000147570), score: 0.85 DR1down-regulator of transcription 1, TBP-binding (negative cofactor 2) (ENSG00000117505), score: 0.7 DTLdenticleless homolog (Drosophila) (ENSG00000143476), score: 0.8 DYDC1DPY30 domain containing 1 (ENSG00000170788), score: 0.78 EFCAB5EF-hand calcium binding domain 5 (ENSG00000176927), score: 0.7 ERCC8excision repair cross-complementing rodent repair deficiency, complementation group 8 (ENSG00000049167), score: 0.7 EYA4eyes absent homolog 4 (Drosophila) (ENSG00000112319), score: 0.7 FAM161Afamily with sequence similarity 161, member A (ENSG00000170264), score: 0.8 FAM194Afamily with sequence similarity 194, member A (ENSG00000163645), score: 0.79 FAM46Cfamily with sequence similarity 46, member C (ENSG00000183508), score: 0.72 FAM48Afamily with sequence similarity 48, member A (ENSG00000102710), score: 0.81 FAM81Bfamily with sequence similarity 81, member B (ENSG00000153347), score: 0.74 FBXO43F-box protein 43 (ENSG00000156509), score: 0.75 FEM1Cfem-1 homolog c (C. elegans) (ENSG00000145780), score: 0.78 FXR1fragile X mental retardation, autosomal homolog 1 (ENSG00000114416), score: 0.68 GKAP1G kinase anchoring protein 1 (ENSG00000165113), score: 0.78 GMPSguanine monphosphate synthetase (ENSG00000163655), score: 0.8 GPR55G protein-coupled receptor 55 (ENSG00000135898), score: 0.73 GRB2growth factor receptor-bound protein 2 (ENSG00000177885), score: -0.77 GSG1germ cell associated 1 (ENSG00000111305), score: 1 HIAT1hippocampus abundant transcript 1 (ENSG00000156875), score: 0.8 HORMAD1HORMA domain containing 1 (ENSG00000143452), score: 0.68 HSF2BPheat shock transcription factor 2 binding protein (ENSG00000160207), score: 0.77 IFT88intraflagellar transport 88 homolog (Chlamydomonas) (ENSG00000032742), score: 0.73 IQCHIQ motif containing H (ENSG00000103599), score: 0.78 ISG20L2interferon stimulated exonuclease gene 20kDa-like 2 (ENSG00000143319), score: 0.72 KIAA0586KIAA0586 (ENSG00000100578), score: 0.77 KIF11kinesin family member 11 (ENSG00000138160), score: 0.72 KIF15kinesin family member 15 (ENSG00000163808), score: 0.72 KIF18Akinesin family member 18A (ENSG00000121621), score: 0.8 KIF27kinesin family member 27 (ENSG00000165115), score: 0.73 KIF2Bkinesin family member 2B (ENSG00000141200), score: 0.71 KLHL10kelch-like 10 (Drosophila) (ENSG00000161594), score: 0.7 KNTC1kinetochore associated 1 (ENSG00000184445), score: 0.77 LASS3LAG1 homolog, ceramide synthase 3 (ENSG00000154227), score: 0.98 LETM2leucine zipper-EF-hand containing transmembrane protein 2 (ENSG00000165046), score: 0.72 LGSNlengsin, lens protein with glutamine synthetase domain (ENSG00000146166), score: 0.74 LIG3ligase III, DNA, ATP-dependent (ENSG00000005156), score: 0.73 LMNB2lamin B2 (ENSG00000176619), score: 0.7 LOC81691exonuclease NEF-sp (ENSG00000005189), score: 0.79 LPPLIM domain containing preferred translocation partner in lipoma (ENSG00000145012), score: 0.7 LRRC43leucine rich repeat containing 43 (ENSG00000158113), score: 0.72 LRRC52leucine rich repeat containing 52 (ENSG00000162763), score: 0.68 LRRC67leucine rich repeat containing 67 (ENSG00000178125), score: 0.72 LRRCC1leucine rich repeat and coiled-coil domain containing 1 (ENSG00000133739), score: 0.87 LRRIQ1leucine-rich repeats and IQ motif containing 1 (ENSG00000133640), score: 0.69 LRRIQ4leucine-rich repeats and IQ motif containing 4 (ENSG00000188306), score: 0.72 MAELmaelstrom homolog (Drosophila) (ENSG00000143194), score: 0.74 MARCH11membrane-associated ring finger (C3HC4) 11 (ENSG00000183654), score: 0.77 MC2Rmelanocortin 2 receptor (adrenocorticotropic hormone) (ENSG00000185231), score: 0.8 MDH1Bmalate dehydrogenase 1B, NAD (soluble) (ENSG00000138400), score: 0.69 MGAT4Cmannosyl (alpha-1,3-)-glycoprotein beta-1,4-N-acetylglucosaminyltransferase, isozyme C (putative) (ENSG00000182050), score: 0.71 MLF2myeloid leukemia factor 2 (ENSG00000089693), score: -0.69 MORN3MORN repeat containing 3 (ENSG00000139714), score: 0.74 MPP1membrane protein, palmitoylated 1, 55kDa (ENSG00000130830), score: -0.73 MYO15Amyosin XVA (ENSG00000091536), score: 0.73 NCBP1nuclear cap binding protein subunit 1, 80kDa (ENSG00000136937), score: 0.73 NEK2NIMA (never in mitosis gene a)-related kinase 2 (ENSG00000117650), score: 0.79 NFIBnuclear factor I/B (ENSG00000147862), score: -0.72 NGLY1N-glycanase 1 (ENSG00000151092), score: 0.71 NRD1nardilysin (N-arginine dibasic convertase) (ENSG00000078618), score: 0.73 NSUN4NOP2/Sun domain family, member 4 (ENSG00000117481), score: 0.78 NUP153nucleoporin 153kDa (ENSG00000124789), score: 0.77 PFN4profilin family, member 4 (ENSG00000176732), score: 0.7 PIAS4protein inhibitor of activated STAT, 4 (ENSG00000105229), score: 0.77 PIWIL1piwi-like 1 (Drosophila) (ENSG00000125207), score: 0.72 POLKpolymerase (DNA directed) kappa (ENSG00000122008), score: 0.79 PPM1Gprotein phosphatase, Mg2+/Mn2+ dependent, 1G (ENSG00000115241), score: 0.86 RAE1RAE1 RNA export 1 homolog (S. pombe) (ENSG00000101146), score: 0.69 RAGErenal tumor antigen (ENSG00000080823), score: 0.67 RANBP9RAN binding protein 9 (ENSG00000010017), score: 0.71 RASEFRAS and EF-hand domain containing (ENSG00000165105), score: 0.7 RBM27RNA binding motif protein 27 (ENSG00000091009), score: 0.67 RFX2regulatory factor X, 2 (influences HLA class II expression) (ENSG00000087903), score: 0.77 RNF17ring finger protein 17 (ENSG00000132972), score: 0.74 RNF32ring finger protein 32 (ENSG00000105982), score: 0.78 RNF38ring finger protein 38 (ENSG00000137075), score: 0.7 ROPN1Lropporin 1-like (ENSG00000145491), score: 0.73 RPS6KA3ribosomal protein S6 kinase, 90kDa, polypeptide 3 (ENSG00000177189), score: -0.7 SCARB2scavenger receptor class B, member 2 (ENSG00000138760), score: -0.73 SHCBP1SHC SH2-domain binding protein 1 (ENSG00000171241), score: 0.79 SLC26A8solute carrier family 26, member 8 (ENSG00000112053), score: 0.71 SLC38A9solute carrier family 38, member 9 (ENSG00000177058), score: 0.78 SPACA1sperm acrosome associated 1 (ENSG00000118434), score: 0.79 SPATA17spermatogenesis associated 17 (ENSG00000162814), score: 0.73 SPATA18spermatogenesis associated 18 homolog (rat) (ENSG00000163071), score: 0.74 SPATA4spermatogenesis associated 4 (ENSG00000150628), score: 0.76 SPO11SPO11 meiotic protein covalently bound to DSB homolog (S. cerevisiae) (ENSG00000054796), score: 0.7 SYCP1synaptonemal complex protein 1 (ENSG00000198765), score: 0.76 TAS1R1taste receptor, type 1, member 1 (ENSG00000173662), score: 0.67 TDP1tyrosyl-DNA phosphodiesterase 1 (ENSG00000042088), score: 0.73 TDRD6tudor domain containing 6 (ENSG00000180113), score: 0.68 TEKT3tektin 3 (ENSG00000125409), score: 0.68 TMC1transmembrane channel-like 1 (ENSG00000165091), score: 0.74 TMC5transmembrane channel-like 5 (ENSG00000103534), score: 0.83 TMF1TATA element modulatory factor 1 (ENSG00000144747), score: 0.76 TRIM24tripartite motif-containing 24 (ENSG00000122779), score: 0.76 TRIM36tripartite motif-containing 36 (ENSG00000152503), score: 0.68 TRIM8tripartite motif-containing 8 (ENSG00000171206), score: -0.78 TRIP12thyroid hormone receptor interactor 12 (ENSG00000153827), score: 0.78 TTC16tetratricopeptide repeat domain 16 (ENSG00000167094), score: 0.78 TTC25tetratricopeptide repeat domain 25 (ENSG00000204815), score: 0.67 TTC29tetratricopeptide repeat domain 29 (ENSG00000137473), score: 0.77 TTKTTK protein kinase (ENSG00000112742), score: 0.72 TTLL9tubulin tyrosine ligase-like family, member 9 (ENSG00000131044), score: 0.73 USP7ubiquitin specific peptidase 7 (herpes virus-associated) (ENSG00000187555), score: 0.76 VAMP7vesicle-associated membrane protein 7 (ENSG00000124333), score: -0.71 VPRBPVpr (HIV-1) binding protein (ENSG00000145041), score: 0.73 VPS13Avacuolar protein sorting 13 homolog A (S. cerevisiae) (ENSG00000197969), score: 0.7 WDR64WD repeat domain 64 (ENSG00000162843), score: 0.74 WDR81WD repeat domain 81 (ENSG00000167716), score: -0.71 ZBTB44zinc finger and BTB domain containing 44 (ENSG00000196323), score: 0.68 ZNF318zinc finger protein 318 (ENSG00000171467), score: 0.71 ZNRF4zinc and ring finger 4 (ENSG00000105428), score: 0.75 ZPBPzona pellucida binding protein (ENSG00000042813), score: 0.75 ZPBP2zona pellucida binding protein 2 (ENSG00000186075), score: 0.75
| Id | species | tissue | sex | individual |
|---|---|---|---|---|
| ppa_ts_m_ca1 | ppa | ts | m | _ |
| ptr_ts_m_ca1 | ptr | ts | m | _ |
| mml_ts_m1_ca1 | mml | ts | m | 1 |
| mml_ts_m2_ca1 | mml | ts | m | 2 |
| mmu_ts_m1_ca1 | mmu | ts | m | 1 |
| mmu_ts_m2_ca1 | mmu | ts | m | 2 |