Overview of the group
The Computational Phylogenetics group is part of the Department of Computational Biology from the University of Lausanne.

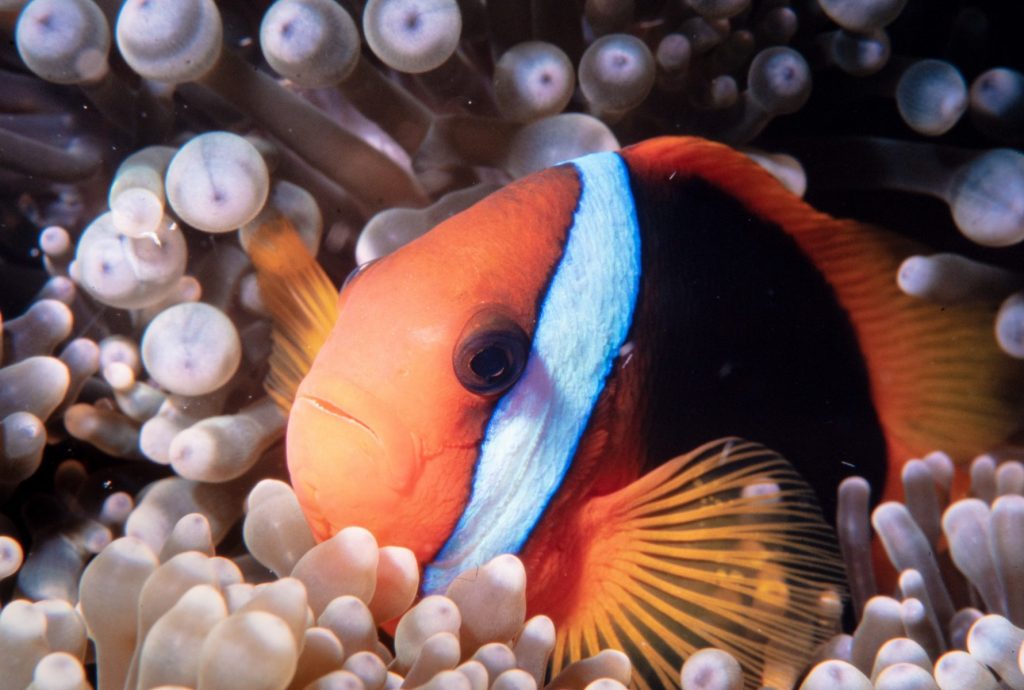
In our group, we work on a variety of organisms and questions with a particular focus on understanding the drivers of adaptation that are associated with the diversification of species. Our current model system is the clownfish adaptive radiation, which is a fascinating group of coral reef fishes that have evolved a mutualistic association with see anemone. We are gathering genomic data for this group to investigate the mechanisms that have driven their evolution and diversification.
Our group is also heavily involved in the development of models of trait evolution, with a particular interest in integrating phenotypic variation in the inference of models of macroevolution. Although our research questions are purely biological and we work on a variety of biological systems, we make extensive use of high performance computing facilities to create and evaluate evolutionary models.
The group is therefore a nice mix of theoretical people, bioinformaticians and empirical biologists, which makes discussions and lab meeting lively and interesting!