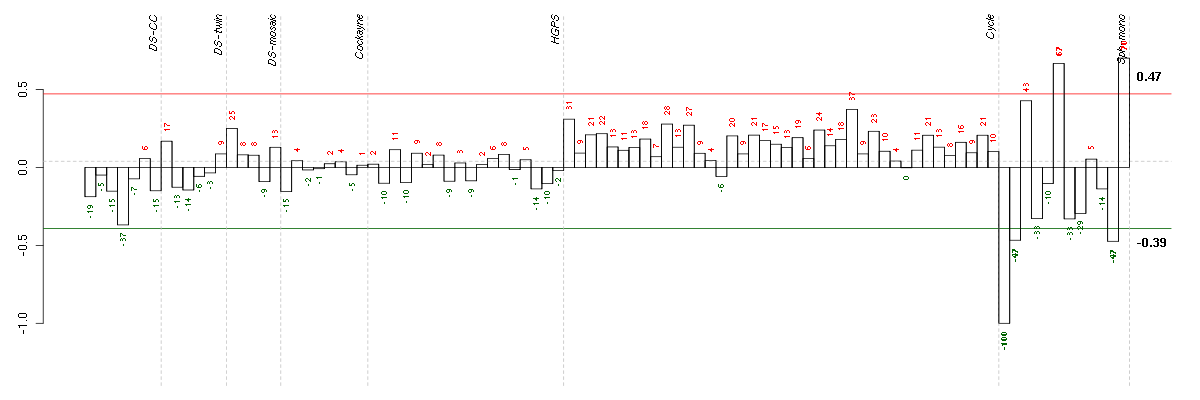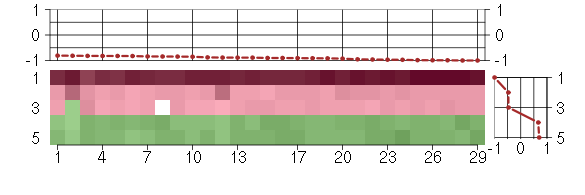

Under-expression is coded with green,
over-expression with red color.



ADORA2Aadenosine A2a receptor (205013_s_at), score: -0.81 ALMS1Alstrom syndrome 1 (214707_x_at), score: -0.92 APBB2amyloid beta (A4) precursor protein-binding, family B, member 2 (212972_x_at), score: -0.99 CDKN1Ccyclin-dependent kinase inhibitor 1C (p57, Kip2) (213183_s_at), score: -0.84 DAPP1dual adaptor of phosphotyrosine and 3-phosphoinositides (219290_x_at), score: -0.91 DNAH3dynein, axonemal, heavy chain 3 (220725_x_at), score: -0.97 FBXW12F-box and WD repeat domain containing 12 (215600_x_at), score: -0.84 FLJ11292hypothetical protein FLJ11292 (220828_s_at), score: -1 G3BP1GTPase activating protein (SH3 domain) binding protein 1 (222187_x_at), score: -0.99 HCG2P7HLA complex group 2 pseudogene 7 (216229_x_at), score: -0.89 IBD12Inflammatory bowel disease 12 (215373_x_at), score: -0.85 LOC643313similar to hypothetical protein LOC284701 (211050_x_at), score: -0.9 LOC647070hypothetical LOC647070 (215467_x_at), score: -0.9 MAP2K7mitogen-activated protein kinase kinase 7 (216206_x_at), score: -0.82 METAP2methionyl aminopeptidase 2 (202015_x_at), score: -0.83 METTL7Amethyltransferase like 7A (211424_x_at), score: -0.89 POM121L9PPOM121 membrane glycoprotein-like 9 (rat) pseudogene (222253_s_at), score: -0.82 SH3GL3SH3-domain GRB2-like 3 (211565_at), score: -0.96 SLC30A5solute carrier family 30 (zinc transporter), member 5 (220181_x_at), score: -0.95 SPINLW1serine peptidase inhibitor-like, with Kunitz and WAP domains 1 (eppin) (206318_at), score: -0.93 SPNsialophorin (206057_x_at), score: -0.99 SPTLC3serine palmitoyltransferase, long chain base subunit 3 (220456_at), score: -0.97 TCTN2tectonic family member 2 (206438_x_at), score: -0.82 TIMM8Atranslocase of inner mitochondrial membrane 8 homolog A (yeast) (210800_at), score: -1 TRIM36tripartite motif-containing 36 (219736_at), score: -0.89 UBQLN4ubiquilin 4 (222252_x_at), score: -0.85 ZC3H7Bzinc finger CCCH-type containing 7B (206169_x_at), score: -0.91 ZNF528zinc finger protein 528 (215019_x_at), score: -0.81 ZNF816Azinc finger protein 816A (217541_x_at), score: -0.88
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-GEOD-4219-raw-cel-1311956083.cel | 2 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956634.cel | 19 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956138.cel | 4 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956358.cel | 10 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| E-GEOD-4219-raw-cel-1311956824.cel | 24 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |