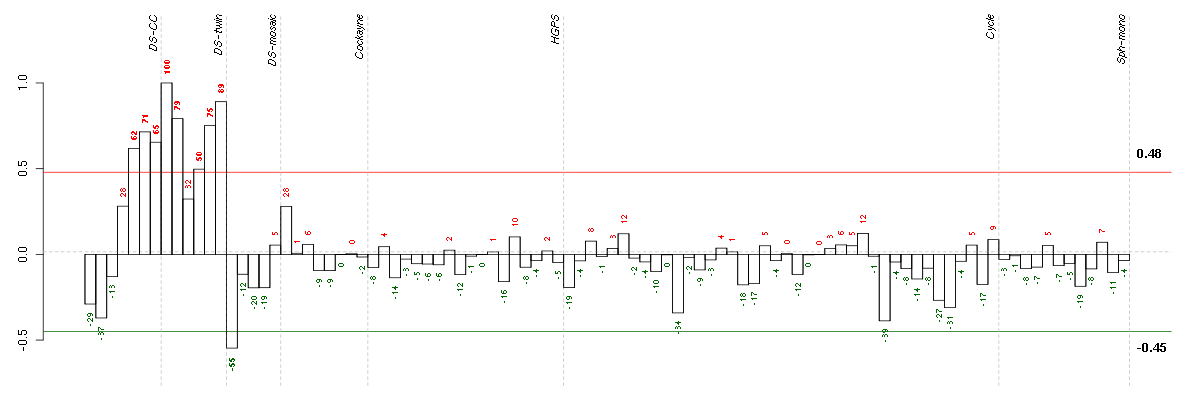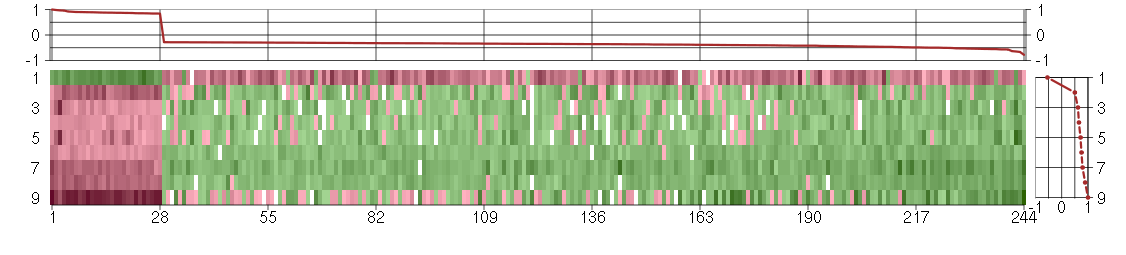

Under-expression is coded with green,
over-expression with red color.

angiogenesis
Blood vessel formation when new vessels emerge from the proliferation of pre-existing blood vessels.
blood vessel development
The process whose specific outcome is the progression of the blood vessel over time, from its formation to the mature structure. The blood vessel is the vasculature carrying blood.
vasculature development
The process whose specific outcome is the progression of the vasculature over time, from its formation to the mature structure.
immune system process
Any process involved in the development or functioning of the immune system, an organismal system for calibrated responses to potential internal or invasive threats.
antigen processing and presentation of peptide antigen via MHC class I
The process by which an antigen-presenting cell expresses a peptide antigen on its cell surface in association with an MHC class I protein complex. Class I here refers to classical class I molecules.
defense response
Reactions, triggered in response to the presence of a foreign body or the occurrence of an injury, which result in restriction of damage to the organism attacked or prevention/recovery from the infection caused by the attack.
signal transduction
The cascade of processes by which a signal interacts with a receptor, causing a change in the level or activity of a second messenger or other downstream target, and ultimately effecting a change in the functioning of the cell.
cell motion
Any process involved in the controlled movement of a cell.
chemotaxis
The directed movement of a motile cell or organism, or the directed growth of a cell guided by a specific chemical concentration gradient. Movement may be towards a higher concentration (positive chemotaxis) or towards a lower concentration (negative chemotaxis).
response to stress
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a disturbance in organismal or cellular homeostasis, usually, but not necessarily, exogenous (e.g. temperature, humidity, ionizing radiation).
inflammatory response
The immediate defensive reaction (by vertebrate tissue) to infection or injury caused by chemical or physical agents. The process is characterized by local vasodilation, extravasation of plasma into intercellular spaces and accumulation of white blood cells and macrophages.
immune response
Any immune system process that functions in the calibrated response of an organism to a potential internal or invasive threat.
cell communication
Any process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
cell surface receptor linked signal transduction
Any series of molecular signals initiated by the binding of an extracellular ligand to a receptor on the surface of the target cell.
enzyme linked receptor protein signaling pathway
Any series of molecular signals initiated by the binding of an extracellular ligand to a receptor on the surface of the target cell, where the receptor possesses catalytic activity or is closely associated with an enzyme such as a protein kinase.
transmembrane receptor protein tyrosine kinase signaling pathway
The series of molecular signals generated as a consequence of a transmembrane receptor tyrosine kinase binding to its physiological ligand.
intracellular signaling cascade
A series of reactions within the cell that occur as a result of a single trigger reaction or compound.
protein kinase cascade
A series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
multicellular organismal development
The biological process whose specific outcome is the progression of a multicellular organism over time from an initial condition (e.g. a zygote or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
anatomical structure morphogenesis
The process by which anatomical structures are generated and organized. Morphogenesis pertains to the creation of form.
organ morphogenesis
Morphogenesis of an organ. An organ is defined as a tissue or set of tissues that work together to perform a specific function or functions. Morphogenesis is the process by which anatomical structures are generated and organized. Organs are commonly observed as visibly distinct structures, but may also exist as loosely associated clusters of cells that work together to perform a specific function or functions.
behavior
The specific actions or reactions of an organism in response to external or internal stimuli. Patterned activity of a whole organism in a manner dependent upon some combination of that organism's internal state and external conditions.
locomotory behavior
The specific movement from place to place of an organism in response to external or internal stimuli. Locomotion of a whole organism in a manner dependent upon some combination of that organism's internal state and external conditions.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
cell proliferation
The multiplication or reproduction of cells, resulting in the expansion of a cell population.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
response to external stimulus
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of an external stimulus.
response to wounding
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a stimulus indicating damage to the organism.
regulation of signal transduction
Any process that modulates the frequency, rate or extent of signal transduction.
positive regulation of signal transduction
Any process that activates or increases the frequency, rate or extent of signal transduction.
cellular process
Any process that is carried out at the cellular level, but not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of protein kinase cascade
Any process that modulates the rate, frequency or extent of a series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
regulation of cell communication
Any process that modulates the frequency, rate or extent of cell communication. Cell communication is the process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
positive regulation of cell communication
Any process that increases the frequency, rate or extent of cell communication. Cell communication is the process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
positive regulation of protein kinase cascade
Any process that increases the rate, frequency or extent of a series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
muscle cell migration
The orderly movement of muscle cells from one site to another, often during the development of a multicellular organism.
cell migration
The orderly movement of cells from one site to another, often during the development of a multicellular organism or multicellular structure.
antigen processing and presentation
The process by which an antigen-presenting cell expresses antigen (peptide or lipid) on its cell surface in association with an MHC protein complex.
cell differentiation
The process whereby relatively unspecialized cells, e.g. embryonic or regenerative cells, acquire specialized structural and/or functional features that characterize the cells, tissues, or organs of the mature organism or some other relatively stable phase of the organism's life history. Differentiation includes the processes involved in commitment of a cell to a specific fate.
regulation of cell migration
Any process that modulates the frequency, rate or extent of cell migration.
leukocyte chemotaxis
The movement of a leukocyte in response to an external stimulus.
multicellular organismal process
Any biological process, occurring at the level of a multicellular organism, pertinent to its function.
developmental process
A biological process whose specific outcome is the progression of an integrated living unit: an anatomical structure (which may be a subcellular structure, cell, tissue, or organ), or organism over time from an initial condition to a later condition.
regulation of localization
Any process that modulates the frequency, rate or extent of any process by which a cell, a substance, or a cellular entity is transported to, or maintained in, a specific location.
locomotion
Self-propelled movement of a cell or organism from one location to another.
regulation of locomotion
Any process that modulates the frequency, rate or extent of locomotion of a cell or organism.
wound healing
The series of events that restore integrity to a damaged tissue, following an injury.
regulation of cell proliferation
Any process that modulates the frequency, rate or extent of cell proliferation.
response to chemical stimulus
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a chemical stimulus.
taxis
The directed movement of a motile cell or organism in response to an external stimulus.
regulation of cell differentiation
Any process that modulates the frequency, rate or extent of cell differentiation, the process whereby relatively unspecialized cells acquire specialized structural and functional features.
positive regulation of cell differentiation
Any process that activates or increases the frequency, rate or extent of cell differentiation.
antigen processing and presentation of peptide antigen
The process by which an antigen-presenting cell expresses peptide antigen in association with an MHC protein complex on its cell surface, including proteolysis and transport steps for the peptide antigen both prior to and following assembly with the MHC protein complex. The peptide antigen is typically, but not always, processed from an endogenous or exogenous protein.
organ development
Development of a tissue or tissues that work together to perform a specific function or functions. Development pertains to the process whose specific outcome is the progression of a structure over time, from its formation to the mature structure. Organs are commonly observed as visibly distinct structures, but may also exist as loosely associated clusters of cells that work together to perform a specific function or functions.
blood vessel morphogenesis
The process by which the anatomical structures of blood vessels are generated and organized. Morphogenesis pertains to the creation of form. The blood vessel is the vasculature carrying blood.
positive regulation of biological process
Any process that activates or increases the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
anatomical structure formation
The process pertaining to the initial formation of an anatomical structure from unspecified parts. This process begins with the specific processes that contribute to the appearance of the discrete structure and ends when the structural rudiment is recognizable. An anatomical structure is any biological entity that occupies space and is distinguished from its surroundings. Anatomical structures can be macroscopic such as a carpel, or microscopic such as an acrosome.
system development
The process whose specific outcome is the progression of an organismal system over time, from its formation to the mature structure. A system is a regularly interacting or interdependent group of organs or tissues that work together to carry out a given biological process.
anatomical structure development
The biological process whose specific outcome is the progression of an anatomical structure from an initial condition to its mature state. This process begins with the formation of the structure and ends with the mature structure, whatever form that may be including its natural destruction. An anatomical structure is any biological entity that occupies space and is distinguished from its surroundings. Anatomical structures can be macroscopic such as a carpel, or microscopic such as an acrosome.
cellular developmental process
A biological process whose specific outcome is the progression of a cell over time from an initial condition to a later condition.
cell motility
Any process involved in the controlled movement of a cell that results in translocation of the cell from one place to another.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of developmental process
Any process that modulates the frequency, rate or extent of development, the biological process whose specific outcome is the progression of a multicellular organism over time from an initial condition (e.g. a zygote, or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
response to stimulus
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a stimulus.
leukocyte migration
The movement of leukocytes within or between different tissues and organs of the body.
positive regulation of developmental process
Any process that activates or increases the rate or extent of development, the biological process whose specific outcome is the progression of an organism over time from an initial condition (e.g. a zygote, or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
localization
Any process by which a cell, a substance, or a cellular entity, such as a protein complex or organelle, is transported to, and/or maintained in a specific location.
regulation of multicellular organismal process
Any process that modulates the frequency, rate or extent of a multicellular organismal process, the processes pertinent to the function of a multicellular organism above the cellular level; includes the integrated processes of tissues and organs.
regulation of cell motion
Any process that modulates the frequency, rate or extent of the movement of a cell.
localization of cell
Any process by which a cell is transported to, and/or maintained in, a specific location.
cell chemotaxis
The directed movement of a motile cell guided by a specific chemical concentration gradient. Movement may be towards a higher concentration (positive chemotaxis) or towards a lower concentration (negative chemotaxis).
biological regulation
Any process that modulates the frequency, rate or extent of any biological process, quality or function.
all
This term is the most general term possible
multicellular organismal development
The biological process whose specific outcome is the progression of a multicellular organism over time from an initial condition (e.g. a zygote or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
cellular developmental process
A biological process whose specific outcome is the progression of a cell over time from an initial condition to a later condition.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
positive regulation of developmental process
Any process that activates or increases the rate or extent of development, the biological process whose specific outcome is the progression of an organism over time from an initial condition (e.g. a zygote, or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
regulation of locomotion
Any process that modulates the frequency, rate or extent of locomotion of a cell or organism.
positive regulation of biological process
Any process that activates or increases the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of developmental process
Any process that modulates the frequency, rate or extent of development, the biological process whose specific outcome is the progression of a multicellular organism over time from an initial condition (e.g. a zygote, or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of multicellular organismal process
Any process that modulates the frequency, rate or extent of a multicellular organismal process, the processes pertinent to the function of a multicellular organism above the cellular level; includes the integrated processes of tissues and organs.
immune response
Any immune system process that functions in the calibrated response of an organism to a potential internal or invasive threat.
regulation of localization
Any process that modulates the frequency, rate or extent of any process by which a cell, a substance, or a cellular entity is transported to, or maintained in, a specific location.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
cell motility
Any process involved in the controlled movement of a cell that results in translocation of the cell from one place to another.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
positive regulation of cell communication
Any process that increases the frequency, rate or extent of cell communication. Cell communication is the process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
signal transduction
The cascade of processes by which a signal interacts with a receptor, causing a change in the level or activity of a second messenger or other downstream target, and ultimately effecting a change in the functioning of the cell.
regulation of cell communication
Any process that modulates the frequency, rate or extent of cell communication. Cell communication is the process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
regulation of cell proliferation
Any process that modulates the frequency, rate or extent of cell proliferation.
positive regulation of cellular process
Any process that activates or increases the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of cell motion
Any process that modulates the frequency, rate or extent of the movement of a cell.
anatomical structure formation
The process pertaining to the initial formation of an anatomical structure from unspecified parts. This process begins with the specific processes that contribute to the appearance of the discrete structure and ends when the structural rudiment is recognizable. An anatomical structure is any biological entity that occupies space and is distinguished from its surroundings. Anatomical structures can be macroscopic such as a carpel, or microscopic such as an acrosome.
anatomical structure morphogenesis
The process by which anatomical structures are generated and organized. Morphogenesis pertains to the creation of form.
system development
The process whose specific outcome is the progression of an organismal system over time, from its formation to the mature structure. A system is a regularly interacting or interdependent group of organs or tissues that work together to carry out a given biological process.
regulation of cell differentiation
Any process that modulates the frequency, rate or extent of cell differentiation, the process whereby relatively unspecialized cells acquire specialized structural and functional features.
positive regulation of developmental process
Any process that activates or increases the rate or extent of development, the biological process whose specific outcome is the progression of an organism over time from an initial condition (e.g. a zygote, or a young adult) to a later condition (e.g. a multicellular animal or an aged adult).
positive regulation of cell differentiation
Any process that activates or increases the frequency, rate or extent of cell differentiation.
regulation of cell motion
Any process that modulates the frequency, rate or extent of the movement of a cell.
response to wounding
A change in state or activity of a cell or an organism (in terms of movement, secretion, enzyme production, gene expression, etc.) as a result of a stimulus indicating damage to the organism.
taxis
The directed movement of a motile cell or organism in response to an external stimulus.
chemotaxis
The directed movement of a motile cell or organism, or the directed growth of a cell guided by a specific chemical concentration gradient. Movement may be towards a higher concentration (positive chemotaxis) or towards a lower concentration (negative chemotaxis).
cell motion
Any process involved in the controlled movement of a cell.
regulation of cell migration
Any process that modulates the frequency, rate or extent of cell migration.
regulation of signal transduction
Any process that modulates the frequency, rate or extent of signal transduction.
positive regulation of cell communication
Any process that increases the frequency, rate or extent of cell communication. Cell communication is the process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
positive regulation of signal transduction
Any process that activates or increases the frequency, rate or extent of signal transduction.
positive regulation of cell proliferation
Any process that activates or increases the rate or extent of cell proliferation.
regulation of cell differentiation
Any process that modulates the frequency, rate or extent of cell differentiation, the process whereby relatively unspecialized cells acquire specialized structural and functional features.
positive regulation of cell differentiation
Any process that activates or increases the frequency, rate or extent of cell differentiation.
positive regulation of cell differentiation
Any process that activates or increases the frequency, rate or extent of cell differentiation.
organ development
Development of a tissue or tissues that work together to perform a specific function or functions. Development pertains to the process whose specific outcome is the progression of a structure over time, from its formation to the mature structure. Organs are commonly observed as visibly distinct structures, but may also exist as loosely associated clusters of cells that work together to perform a specific function or functions.
organ morphogenesis
Morphogenesis of an organ. An organ is defined as a tissue or set of tissues that work together to perform a specific function or functions. Morphogenesis is the process by which anatomical structures are generated and organized. Organs are commonly observed as visibly distinct structures, but may also exist as loosely associated clusters of cells that work together to perform a specific function or functions.
cell chemotaxis
The directed movement of a motile cell guided by a specific chemical concentration gradient. Movement may be towards a higher concentration (positive chemotaxis) or towards a lower concentration (negative chemotaxis).
regulation of cell migration
Any process that modulates the frequency, rate or extent of cell migration.
leukocyte migration
The movement of leukocytes within or between different tissues and organs of the body.
leukocyte chemotaxis
The movement of a leukocyte in response to an external stimulus.
inflammatory response
The immediate defensive reaction (by vertebrate tissue) to infection or injury caused by chemical or physical agents. The process is characterized by local vasodilation, extravasation of plasma into intercellular spaces and accumulation of white blood cells and macrophages.
taxis
The directed movement of a motile cell or organism in response to an external stimulus.
positive regulation of signal transduction
Any process that activates or increases the frequency, rate or extent of signal transduction.
angiogenesis
Blood vessel formation when new vessels emerge from the proliferation of pre-existing blood vessels.
regulation of protein kinase cascade
Any process that modulates the rate, frequency or extent of a series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
positive regulation of protein kinase cascade
Any process that increases the rate, frequency or extent of a series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
positive regulation of protein kinase cascade
Any process that increases the rate, frequency or extent of a series of reactions, mediated by protein kinases, which occurs as a result of a single trigger reaction or compound.
blood vessel morphogenesis
The process by which the anatomical structures of blood vessels are generated and organized. Morphogenesis pertains to the creation of form. The blood vessel is the vasculature carrying blood.

plasma membrane
The membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
membrane
Double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
MHC class I protein complex
A transmembrane protein complex composed of a MHC class I alpha chain and an invariant beta2-microglobin chain, and with or without a bound peptide antigen. Class I here refers to classical class I molecules.
integral to membrane
Penetrating at least one phospholipid bilayer of a membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer. When used to describe a protein, indicates that all or part of the peptide sequence is embedded in the membrane.
extracellular region
The space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers the host cell environment outside an intracellular parasite.
integral to plasma membrane
Penetrating at least one phospholipid bilayer of a plasma membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
extracellular space
That part of a multicellular organism outside the cells proper, usually taken to be outside the plasma membranes, and occupied by fluid.
cell
The basic structural and functional unit of all organisms. Includes the plasma membrane and any external encapsulating structures such as the cell wall and cell envelope.
intrinsic to membrane
Located in a membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
intrinsic to plasma membrane
Located in the plasma membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
macromolecular complex
A stable assembly of two or more macromolecules, i.e. proteins, nucleic acids, carbohydrates or lipids, in which the constituent parts function together.
MHC protein complex
A transmembrane protein complex composed of an MHC alpha chain and, in most cases, either an MHC class II beta chain or an invariant beta2-microglobin chain, and with or without a bound peptide, lipid, or polysaccharide antigen.
protein complex
Any macromolecular complex composed of two or more polypeptide subunits, which may or may not be identical. Protein complexes may have other associated non-protein prosthetic groups, such as nucleotides, metal ions or carbohydrate groups.
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
plasma membrane part
Any constituent part of the plasma membrane, the membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
all
This term is the most general term possible
extracellular region part
Any constituent part of the extracellular region, the space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers constituent parts of the host cell environment outside an intracellular parasite.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
plasma membrane part
Any constituent part of the plasma membrane, the membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
intrinsic to plasma membrane
Located in the plasma membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
MHC protein complex
A transmembrane protein complex composed of an MHC alpha chain and, in most cases, either an MHC class II beta chain or an invariant beta2-microglobin chain, and with or without a bound peptide, lipid, or polysaccharide antigen.
integral to plasma membrane
Penetrating at least one phospholipid bilayer of a plasma membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer.

protein binding
Interacting selectively with any protein or protein complex (a complex of two or more proteins that may include other nonprotein molecules).
molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
receptor binding
Interacting selectively with one or more specific sites on a receptor molecule, a macromolecule that undergoes combination with a hormone, neurotransmitter, drug or intracellular messenger to initiate a change in cell function.
signal transducer activity
Mediates the transfer of a signal from the outside to the inside of a cell by means other than the introduction of the signal molecule itself into the cell.
receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity.
transmembrane receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity, and spanning to the membrane of either the cell or an organelle.
cytokine activity
Functions to control the survival, growth, differentiation and effector function of tissues and cells.
binding
The selective, often stoichiometric, interaction of a molecule with one or more specific sites on another molecule.
growth factor activity
The function that stimulates a cell to grow or proliferate. Most growth factors have other actions besides the induction of cell growth or proliferation.
MHC class I receptor activity
Combining with an MHC class I protein complex to initiate a change in cellular activity. Class I here refers to classical class I molecules.
molecular transducer activity
The molecular function that accepts an input of one form and creates an output of a different form.
all
This term is the most general term possible
| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 04060 | 1.311e-03 | 4.107 | 15 | 115 | Cytokine-cytokine receptor interaction |
| 04610 | 1.885e-03 | 1.179 | 8 | 33 | Complement and coagulation cascades |
| 05332 | 6.323e-03 | 0.75 | 6 | 21 | Graft-versus-host disease |
| 04940 | 1.776e-02 | 0.9286 | 6 | 26 | Type I diabetes mellitus |
| 05330 | 3.877e-02 | 0.75 | 5 | 21 | Allograft rejection |
| 05320 | 4.594e-02 | 0.7857 | 5 | 22 | Autoimmune thyroid disease |
| 04630 | 4.614e-02 | 3.107 | 10 | 87 | Jak-STAT signaling pathway |
A2Malpha-2-macroglobulin (217757_at), score: -0.29 ABLIM3actin binding LIM protein family, member 3 (205730_s_at), score: -0.36 ADRA2Aadrenergic, alpha-2A-, receptor (209869_at), score: -0.54 AJAP1adherens junctions associated protein 1 (206460_at), score: -0.44 ANGPTL4angiopoietin-like 4 (221009_s_at), score: -0.45 ANO1anoctamin 1, calcium activated chloride channel (218804_at), score: -0.45 APODapolipoprotein D (201525_at), score: -0.34 ARID5AAT rich interactive domain 5A (MRF1-like) (213138_at), score: -0.31 ASAP2ArfGAP with SH3 domain, ankyrin repeat and PH domain 2 (206414_s_at), score: -0.29 ATP13A3ATPase type 13A3 (219558_at), score: -0.35 B4GALT5UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 5 (221484_at), score: -0.35 B4GALT6UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 6 (206233_at), score: 0.86 BCL7AB-cell CLL/lymphoma 7A (203795_s_at), score: -0.3 BMPR2bone morphogenetic protein receptor, type II (serine/threonine kinase) (210214_s_at), score: -0.32 C17orf91chromosome 17 open reading frame 91 (214696_at), score: -0.29 C19orf28chromosome 19 open reading frame 28 (220178_at), score: -0.3 C1Rcomplement component 1, r subcomponent (212067_s_at), score: -0.46 C1Scomplement component 1, s subcomponent (208747_s_at), score: -0.31 C9orf167chromosome 9 open reading frame 167 (219620_x_at), score: -0.31 CADPS2Ca++-dependent secretion activator 2 (219572_at), score: -0.35 CASP1caspase 1, apoptosis-related cysteine peptidase (interleukin 1, beta, convertase) (211368_s_at), score: -0.35 CASP4caspase 4, apoptosis-related cysteine peptidase (209310_s_at), score: -0.35 CBLBCas-Br-M (murine) ecotropic retroviral transforming sequence b (209682_at), score: -0.29 CCL2chemokine (C-C motif) ligand 2 (216598_s_at), score: -0.35 CCRL1chemokine (C-C motif) receptor-like 1 (220351_at), score: -0.46 CD302CD302 molecule (203799_at), score: -0.46 CD55CD55 molecule, decay accelerating factor for complement (Cromer blood group) (201925_s_at), score: -0.28 CDC42SE1CDC42 small effector 1 (218157_x_at), score: -0.44 CLEC2BC-type lectin domain family 2, member B (209732_at), score: -0.42 CLEC7AC-type lectin domain family 7, member A (221698_s_at), score: 0.88 CLIP1CAP-GLY domain containing linker protein 1 (210716_s_at), score: -0.29 COL15A1collagen, type XV, alpha 1 (203477_at), score: -0.63 CREG1cellular repressor of E1A-stimulated genes 1 (201200_at), score: -0.49 CSF1colony stimulating factor 1 (macrophage) (209716_at), score: -0.37 CSGALNACT2chondroitin sulfate N-acetylgalactosaminyltransferase 2 (222235_s_at), score: -0.33 CST3cystatin C (201360_at), score: -0.37 CTSOcathepsin O (203758_at), score: -0.28 CTTNBP2NLCTTNBP2 N-terminal like (214731_at), score: 0.84 CYP51A1cytochrome P450, family 51, subfamily A, polypeptide 1 (216607_s_at), score: -0.36 DCNdecorin (211896_s_at), score: -0.41 DDX3YDEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked (205000_at), score: -0.41 DGKDdiacylglycerol kinase, delta 130kDa (208072_s_at), score: -0.4 DKK2dickkopf homolog 2 (Xenopus laevis) (219908_at), score: -0.55 DNASE2deoxyribonuclease II, lysosomal (214992_s_at), score: -0.32 DOCK1dedicator of cytokinesis 1 (203187_at), score: -0.3 DOCK9dedicator of cytokinesis 9 (212538_at), score: -0.37 DPP4dipeptidyl-peptidase 4 (211478_s_at), score: -0.53 DUSP14dual specificity phosphatase 14 (203367_at), score: -0.36 DUSP3dual specificity phosphatase 3 (201536_at), score: -0.32 ECM1extracellular matrix protein 1 (209365_s_at), score: -0.47 EDN1endothelin 1 (218995_s_at), score: -0.34 EFHD1EF-hand domain family, member D1 (209343_at), score: -0.33 EGLN2egl nine homolog 2 (C. elegans) (220956_s_at), score: 0.84 EGR3early growth response 3 (206115_at), score: -0.4 EIF1AYeukaryotic translation initiation factor 1A, Y-linked (204409_s_at), score: -0.33 EPHB3EPH receptor B3 (1438_at), score: 0.87 ERAP2endoplasmic reticulum aminopeptidase 2 (219759_at), score: -0.29 EXD3exonuclease 3'-5' domain containing 3 (220838_at), score: 0.89 F10coagulation factor X (205620_at), score: -0.52 F3coagulation factor III (thromboplastin, tissue factor) (204363_at), score: -0.43 FADS3fatty acid desaturase 3 (216080_s_at), score: -0.32 FGF2fibroblast growth factor 2 (basic) (204421_s_at), score: -0.3 FGF5fibroblast growth factor 5 (210310_s_at), score: -0.32 FGF7fibroblast growth factor 7 (keratinocyte growth factor) (205782_at), score: -0.46 FHOD1formin homology 2 domain containing 1 (218530_at), score: -0.37 FLGfilaggrin (215704_at), score: 0.97 FLJ10404hypothetical protein FLJ10404 (218920_at), score: 0.85 FNDC3Afibronectin type III domain containing 3A (202304_at), score: -0.29 FOSBFBJ murine osteosarcoma viral oncogene homolog B (202768_at), score: -0.39 GAP43growth associated protein 43 (204471_at), score: 1 GDF5growth differentiation factor 5 (206614_at), score: 0.9 GEMGTP binding protein overexpressed in skeletal muscle (204472_at), score: -0.34 GLT8D2glycosyltransferase 8 domain containing 2 (221447_s_at), score: -0.29 GNALguanine nucleotide binding protein (G protein), alpha activating activity polypeptide, olfactory type (206355_at), score: 0.85 GPNMBglycoprotein (transmembrane) nmb (201141_at), score: -0.78 GPR183G protein-coupled receptor 183 (205419_at), score: -0.3 GPR37G protein-coupled receptor 37 (endothelin receptor type B-like) (209631_s_at), score: -0.38 GPX3glutathione peroxidase 3 (plasma) (214091_s_at), score: -0.42 GRAMD3GRAM domain containing 3 (218706_s_at), score: -0.35 GRIK2glutamate receptor, ionotropic, kainate 2 (213845_at), score: -0.3 GRNgranulin (211284_s_at), score: -0.4 GSTT1glutathione S-transferase theta 1 (203815_at), score: -0.34 HBEGFheparin-binding EGF-like growth factor (203821_at), score: -0.39 HHEXhematopoietically expressed homeobox (215933_s_at), score: -0.36 HIST1H2AChistone cluster 1, H2ac (215071_s_at), score: -0.3 HIST1H2BKhistone cluster 1, H2bk (209806_at), score: -0.31 HK2hexokinase 2 (202934_at), score: -0.49 HLA-Amajor histocompatibility complex, class I, A (213932_x_at), score: -0.4 HLA-Amajor histocompatibility complex, class I, A (217436_x_at), score: -0.4 HLA-Bmajor histocompatibility complex, class I, B (211911_x_at), score: -0.39 HLA-Cmajor histocompatibility complex, class I, C (211799_x_at), score: -0.53 HLA-Emajor histocompatibility complex, class I, E (200904_at), score: -0.55 HLA-Fmajor histocompatibility complex, class I, F (204806_x_at), score: -0.48 HMOX1heme oxygenase (decycling) 1 (203665_at), score: -0.38 HOXB6homeobox B6 (205366_s_at), score: -0.4 HOXB7homeobox B7 (204779_s_at), score: -0.36 HOXD11homeobox D11 (214604_at), score: 0.96 HPCAL1hippocalcin-like 1 (205462_s_at), score: -0.41 HSPB7heat shock 27kDa protein family, member 7 (cardiovascular) (218934_s_at), score: -0.31 HTR2A5-hydroxytryptamine (serotonin) receptor 2A (207135_at), score: -0.38 HTR2B5-hydroxytryptamine (serotonin) receptor 2B (206638_at), score: -0.43 IDI1isopentenyl-diphosphate delta isomerase 1 (204615_x_at), score: -0.38 IFITM1interferon induced transmembrane protein 1 (9-27) (214022_s_at), score: -0.36 IGFBP5insulin-like growth factor binding protein 5 (203424_s_at), score: -0.28 IL1RAPinterleukin 1 receptor accessory protein (205227_at), score: -0.43 IL1RL1interleukin 1 receptor-like 1 (207526_s_at), score: -0.65 IL23Ainterleukin 23, alpha subunit p19 (211796_s_at), score: -0.31 IL6interleukin 6 (interferon, beta 2) (205207_at), score: -0.54 IL6STinterleukin 6 signal transducer (gp130, oncostatin M receptor) (211000_s_at), score: -0.29 IL8interleukin 8 (211506_s_at), score: -0.43 ITGA1integrin, alpha 1 (214660_at), score: -0.31 ITIH2inter-alpha (globulin) inhibitor H2 (204987_at), score: 0.9 ITPR3inositol 1,4,5-triphosphate receptor, type 3 (201189_s_at), score: -0.36 JARID2jumonji, AT rich interactive domain 2 (203297_s_at), score: -0.34 JHDM1Djumonji C domain containing histone demethylase 1 homolog D (S. cerevisiae) (221778_at), score: -0.3 JMJD3jumonji domain containing 3, histone lysine demethylase (213146_at), score: -0.39 KANK1KN motif and ankyrin repeat domains 1 (213005_s_at), score: -0.34 KCNMB4potassium large conductance calcium-activated channel, subfamily M, beta member 4 (219287_at), score: 0.86 KCNN4potassium intermediate/small conductance calcium-activated channel, subfamily N, member 4 (204401_at), score: -0.33 KHDRBS3KH domain containing, RNA binding, signal transduction associated 3 (209781_s_at), score: -0.44 KIAA1644KIAA1644 (52837_at), score: -0.32 KLHL21kelch-like 21 (Drosophila) (203068_at), score: -0.3 KLHL26kelch-like 26 (Drosophila) (219354_at), score: -0.35 LEPROTleptin receptor overlapping transcript (202378_s_at), score: -0.31 LGR4leucine-rich repeat-containing G protein-coupled receptor 4 (218326_s_at), score: 0.88 LHFPlipoma HMGIC fusion partner (218656_s_at), score: -0.5 LIFleukemia inhibitory factor (cholinergic differentiation factor) (205266_at), score: -0.28 LMBRD1LMBR1 domain containing 1 (218191_s_at), score: -0.5 LMCD1LIM and cysteine-rich domains 1 (218574_s_at), score: -0.4 LOC100128809similar to hCG2045829 (215707_s_at), score: -0.36 LOC286434hypothetical protein LOC286434 (222196_at), score: 0.91 LY96lymphocyte antigen 96 (206584_at), score: -0.47 LYNv-yes-1 Yamaguchi sarcoma viral related oncogene homolog (202625_at), score: -0.31 MAFv-maf musculoaponeurotic fibrosarcoma oncogene homolog (avian) (209348_s_at), score: -0.33 MAGI2membrane associated guanylate kinase, WW and PDZ domain containing 2 (209737_at), score: -0.32 MAN1C1mannosidase, alpha, class 1C, member 1 (218918_at), score: -0.55 MAN2B1mannosidase, alpha, class 2B, member 1 (209166_s_at), score: -0.34 MAP2K3mitogen-activated protein kinase kinase 3 (207667_s_at), score: -0.48 MAP3K7IP2mitogen-activated protein kinase kinase kinase 7 interacting protein 2 (210284_s_at), score: -0.35 MARCH7membrane-associated ring finger (C3HC4) 7 (202654_x_at), score: -0.31 MEIS1Meis homeobox 1 (204069_at), score: -0.33 MEIS3P1Meis homeobox 3 pseudogene 1 (214077_x_at), score: -0.42 MEOX1mesenchyme homeobox 1 (205619_s_at), score: 0.9 MFAP4microfibrillar-associated protein 4 (212713_at), score: -0.36 MTSS1metastasis suppressor 1 (203037_s_at), score: -0.34 MYO1Dmyosin ID (212338_at), score: -0.36 NACAP1nascent-polypeptide-associated complex alpha polypeptide pseudogene 1 (211445_x_at), score: 0.85 NAMPTnicotinamide phosphoribosyltransferase (217738_at), score: -0.39 NFATC1nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 1 (210162_s_at), score: -0.55 NFIBnuclear factor I/B (209290_s_at), score: 0.87 NFIL3nuclear factor, interleukin 3 regulated (203574_at), score: -0.32 NFKB1nuclear factor of kappa light polypeptide gene enhancer in B-cells 1 (209239_at), score: -0.54 NID1nidogen 1 (202008_s_at), score: -0.45 NINJ1ninjurin 1 (203045_at), score: -0.39 NOVA1neuro-oncological ventral antigen 1 (205794_s_at), score: 0.89 NPnucleoside phosphorylase (201695_s_at), score: -0.33 NPC1Niemann-Pick disease, type C1 (202679_at), score: -0.43 NPC2Niemann-Pick disease, type C2 (200701_at), score: -0.51 NR3C1nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) (201866_s_at), score: -0.34 NR4A3nuclear receptor subfamily 4, group A, member 3 (209959_at), score: -0.5 OLFML2Aolfactomedin-like 2A (213075_at), score: -0.35 OSTM1osteopetrosis associated transmembrane protein 1 (218196_at), score: -0.37 PCDH7protocadherin 7 (205534_at), score: -0.3 PDGFAplatelet-derived growth factor alpha polypeptide (205463_s_at), score: -0.38 PDGFDplatelet derived growth factor D (219304_s_at), score: -0.39 PDLIM3PDZ and LIM domain 3 (209621_s_at), score: -0.32 PIAS4protein inhibitor of activated STAT, 4 (212881_at), score: 0.84 PKP2plakophilin 2 (207717_s_at), score: -0.46 PLAURplasminogen activator, urokinase receptor (211924_s_at), score: -0.3 PMEPA1prostate transmembrane protein, androgen induced 1 (217875_s_at), score: -0.57 POLR1Cpolymerase (RNA) I polypeptide C, 30kDa (207515_s_at), score: -0.29 PPAP2Aphosphatidic acid phosphatase type 2A (209147_s_at), score: -0.42 PPARDperoxisome proliferator-activated receptor delta (37152_at), score: -0.46 PSMB8proteasome (prosome, macropain) subunit, beta type, 8 (large multifunctional peptidase 7) (209040_s_at), score: -0.29 PTGDSprostaglandin D2 synthase 21kDa (brain) (212187_x_at), score: -0.49 PTPRNprotein tyrosine phosphatase, receptor type, N (204945_at), score: 0.89 RABGGTBRab geranylgeranyltransferase, beta subunit (209180_at), score: -0.32 RALGPS2Ral GEF with PH domain and SH3 binding motif 2 (220338_at), score: 0.92 RANBP2RAN binding protein 2 (201711_x_at), score: -0.32 RAPGEF2Rap guanine nucleotide exchange factor (GEF) 2 (203097_s_at), score: -0.37 RCAN2regulator of calcineurin 2 (203498_at), score: -0.56 RCL1RNA terminal phosphate cyclase-like 1 (218544_s_at), score: -0.33 RGS2regulator of G-protein signaling 2, 24kDa (202388_at), score: -0.3 RGS3regulator of G-protein signaling 3 (203823_at), score: -0.32 RPS4Y1ribosomal protein S4, Y-linked 1 (201909_at), score: -0.53 RRADRas-related associated with diabetes (204802_at), score: -0.67 RUNX1runt-related transcription factor 1 (209360_s_at), score: -0.37 SCARB1scavenger receptor class B, member 1 (201819_at), score: -0.32 SCARB2scavenger receptor class B, member 2 (201647_s_at), score: -0.32 SCPEP1serine carboxypeptidase 1 (218217_at), score: -0.32 SECTM1secreted and transmembrane 1 (213716_s_at), score: -0.34 SEMA5Asema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5A (205405_at), score: -0.31 SEPP1selenoprotein P, plasma, 1 (201427_s_at), score: -0.35 SERPINB2serpin peptidase inhibitor, clade B (ovalbumin), member 2 (204614_at), score: -0.37 SFRS5splicing factor, arginine/serine-rich 5 (212266_s_at), score: -0.33 SGPP1sphingosine-1-phosphate phosphatase 1 (221268_s_at), score: -0.29 SH2B2SH2B adaptor protein 2 (205367_at), score: 0.88 SHBSrc homology 2 domain containing adaptor protein B (204657_s_at), score: -0.32 SIX1SIX homeobox 1 (205817_at), score: 0.87 SLC19A2solute carrier family 19 (thiamine transporter), member 2 (209681_at), score: -0.36 SLC39A8solute carrier family 39 (zinc transporter), member 8 (209267_s_at), score: -0.35 SLC4A5solute carrier family 4, sodium bicarbonate cotransporter, member 5 (204296_at), score: 0.85 SLCO3A1solute carrier organic anion transporter family, member 3A1 (219229_at), score: -0.32 SOCS2suppressor of cytokine signaling 2 (203373_at), score: -0.3 SOD3superoxide dismutase 3, extracellular (205236_x_at), score: -0.39 SOX11SRY (sex determining region Y)-box 11 (204914_s_at), score: 0.98 SOX4SRY (sex determining region Y)-box 4 (201417_at), score: -0.32 SPRED2sprouty-related, EVH1 domain containing 2 (212458_at), score: -0.35 SPRY2sprouty homolog 2 (Drosophila) (204011_at), score: -0.34 SPRY4sprouty homolog 4 (Drosophila) (221489_s_at), score: -0.42 SQRDLsulfide quinone reductase-like (yeast) (217995_at), score: -0.29 STK38Lserine/threonine kinase 38 like (212572_at), score: -0.38 STOMstomatin (201060_x_at), score: -0.41 SVEP1sushi, von Willebrand factor type A, EGF and pentraxin domain containing 1 (213247_at), score: -0.47 TBX3T-box 3 (219682_s_at), score: -0.42 TEX2testis expressed 2 (218099_at), score: -0.34 THBDthrombomodulin (203887_s_at), score: -0.29 TM2D1TM2 domain containing 1 (211703_s_at), score: -0.5 TM6SF1transmembrane 6 superfamily member 1 (219892_at), score: -0.38 TMBIM4transmembrane BAX inhibitor motif containing 4 (219206_x_at), score: -0.36 TMEM41Btransmembrane protein 41B (212623_at), score: -0.5 TNFAIP2tumor necrosis factor, alpha-induced protein 2 (202510_s_at), score: -0.51 TNFRSF11Btumor necrosis factor receptor superfamily, member 11b (204933_s_at), score: -0.49 TNFRSF1Btumor necrosis factor receptor superfamily, member 1B (203508_at), score: -0.29 TNFSF4tumor necrosis factor (ligand) superfamily, member 4 (207426_s_at), score: -0.39 TNS3tensin 3 (217853_at), score: -0.36 TOR1AIP1torsin A interacting protein 1 (216100_s_at), score: -0.29 TP53BP2tumor protein p53 binding protein, 2 (203120_at), score: -0.37 TP53I11tumor protein p53 inducible protein 11 (214667_s_at), score: -0.4 TPD52tumor protein D52 (201690_s_at), score: -0.29 TPP1tripeptidyl peptidase I (200742_s_at), score: -0.56 TPST1tyrosylprotein sulfotransferase 1 (204140_at), score: -0.44 TPST2tyrosylprotein sulfotransferase 2 (204079_at), score: -0.3 TRAF3IP2TRAF3 interacting protein 2 (215411_s_at), score: -0.34 TRIB1tribbles homolog 1 (Drosophila) (202241_at), score: -0.4 TRIM22tripartite motif-containing 22 (213293_s_at), score: -0.35 UTP3UTP3, small subunit (SSU) processome component, homolog (S. cerevisiae) (209486_at), score: -0.29 VEGFAvascular endothelial growth factor A (211527_x_at), score: -0.39 WTAPWilms tumor 1 associated protein (210285_x_at), score: -0.42 YRDCyrdC domain containing (E. coli) (218647_s_at), score: -0.33 ZKSCAN3zinc finger with KRAB and SCAN domains 3 (211773_s_at), score: 0.88 ZMYM6zinc finger, MYM-type 6 (219924_s_at), score: -0.38 ZNF35zinc finger protein 35 (206096_at), score: -0.35 ZNF672zinc finger protein 672 (218068_s_at), score: -0.33
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| 4Twin.CEL | 4 | 2 | DS-twin | hgu133plus2 | none | DS-twin 4 |
| t21b 08-03.CEL | 5 | 1 | DS-CC | hgu133a | Down | DS-CC 5 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| t21c 08-03.CEL | 6 | 1 | DS-CC | hgu133a | Down | DS-CC 6 |
| 5CTwin.CEL | 5 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 5 |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| 6Twin.CEL | 6 | 2 | DS-twin | hgu133plus2 | none | DS-twin 6 |
| 1Twin.CEL | 1 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 1 |