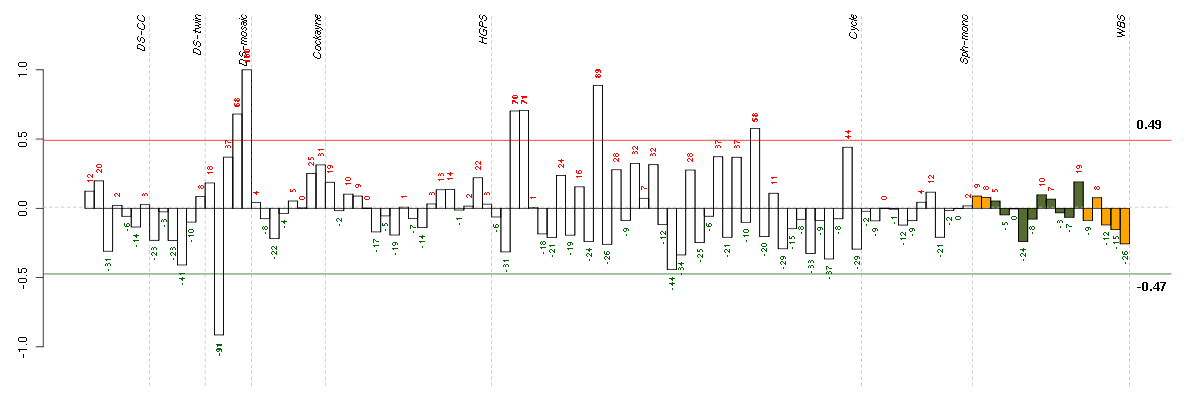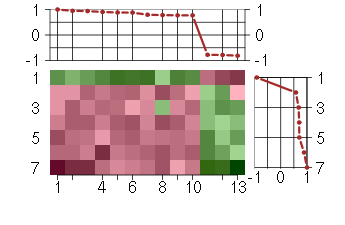

Under-expression is coded with green,
over-expression with red color.



| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 04510 | 1.192e-02 | 0.3368 | 4 | 153 | Focal adhesion |
| 04010 | 2.114e-02 | 0.4094 | 4 | 186 | MAPK signaling pathway |
| 04012 | 2.161e-02 | 0.1519 | 3 | 69 | ErbB signaling pathway |
CFIcomplement factor I (203854_at), score: -0.81 CMAHcytidine monophosphate-N-acetylneuraminic acid hydroxylase (CMP-N-acetylneuraminate monooxygenase) pseudogene (205518_s_at), score: 0.9 EGFepidermal growth factor (beta-urogastrone) (206254_at), score: -0.78 ELK1ELK1, member of ETS oncogene family (210376_x_at), score: 0.77 HMGA1high mobility group AT-hook 1 (210457_x_at), score: 0.77 MACROD1MACRO domain containing 1 (219188_s_at), score: -0.78 MRPS6mitochondrial ribosomal protein S6 (213167_s_at), score: 0.95 PAK1p21 protein (Cdc42/Rac)-activated kinase 1 (209615_s_at), score: 0.78 PXNpaxillin (211823_s_at), score: 0.88 RASA3RAS p21 protein activator 3 (206220_s_at), score: 0.8 SLC29A1solute carrier family 29 (nucleoside transporters), member 1 (201801_s_at), score: 0.94 TGFBR2transforming growth factor, beta receptor II (70/80kDa) (207334_s_at), score: 0.88 TOM1L2target of myb1-like 2 (chicken) (214840_at), score: 1
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| 46B.CEL | 2 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 2 |
| E-TABM-263-raw-cel-1515486211.cel | 29 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 47B.CEL | 4 | 3 | DS-mosaic | hgu133plus2 | Down mosaic | DS-mosaic 4 |
| E-TABM-263-raw-cel-1515485691.cel | 3 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 47C.CEL | 5 | 3 | DS-mosaic | hgu133plus2 | Down mosaic | DS-mosaic 5 |