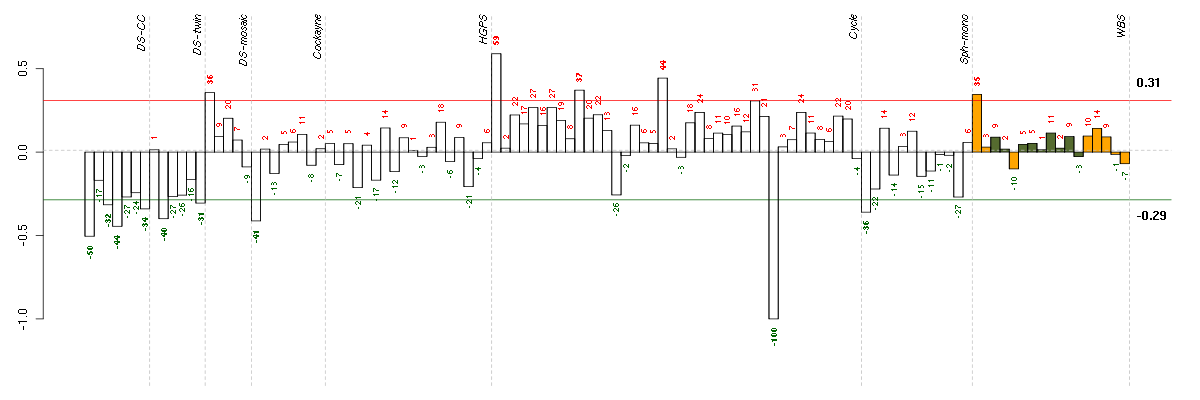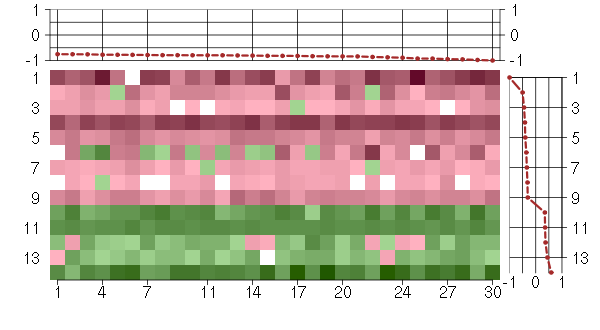

Under-expression is coded with green,
over-expression with red color.



ADRBK2adrenergic, beta, receptor kinase 2 (204184_s_at), score: -0.82 AFF2AF4/FMR2 family, member 2 (206105_at), score: -0.83 ATRNL1attractin-like 1 (213744_at), score: -0.81 C7orf28Achromosome 7 open reading frame 28A (201974_s_at), score: -0.82 CA1carbonic anhydrase I (205949_at), score: -0.92 CD53CD53 molecule (203416_at), score: -0.89 CLDN10claudin 10 (205328_at), score: -0.84 CNKSR2connector enhancer of kinase suppressor of Ras 2 (206731_at), score: -0.96 CTRLchymotrypsin-like (214377_s_at), score: -0.77 DNM3dynamin 3 (209839_at), score: -0.8 GRHL2grainyhead-like 2 (Drosophila) (219388_at), score: -0.93 IL1RAPL1interleukin 1 receptor accessory protein-like 1 (220663_at), score: -0.75 KRT2keratin 2 (207908_at), score: -0.95 LOC93432maltase-glucoamylase-like pseudogene (216666_at), score: -0.8 MSTP9macrophage stimulating, pseudogene 9 (213382_at), score: -0.84 NTRK2neurotrophic tyrosine kinase, receptor, type 2 (207152_at), score: -0.82 PECAM1platelet/endothelial cell adhesion molecule (208982_at), score: -0.78 PEG3paternally expressed 3 (209242_at), score: -0.8 POU4F2POU class 4 homeobox 2 (207725_at), score: -0.86 PPIAL4Apeptidylprolyl isomerase A (cyclophilin A)-like 4A (217136_at), score: -0.78 PROL1proline rich, lacrimal 1 (208004_at), score: -0.78 RAPGEF4Rap guanine nucleotide exchange factor (GEF) 4 (205651_x_at), score: -0.8 RPGRIP1retinitis pigmentosa GTPase regulator interacting protein 1 (206608_s_at), score: -1 SIRPB1signal-regulatory protein beta 1 (206934_at), score: -0.98 SLC26A4solute carrier family 26, member 4 (206529_x_at), score: -0.76 SPAM1sperm adhesion molecule 1 (PH-20 hyaluronidase, zona pellucida binding) (216989_at), score: -0.76 TAAR2trace amine associated receptor 2 (221394_at), score: -0.79 TACR1tachykinin receptor 1 (208048_at), score: -0.88 TP63tumor protein p63 (209863_s_at), score: -0.84 ZNF335zinc finger protein 335 (78330_at), score: -0.79
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515486251.cel | 31 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| E-GEOD-4219-raw-cel-1311956083.cel | 2 | 7 | Sph-mono | hgu133plus2 | none | Sph-mon 1 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| ctrl c 08-03.CEL | 3 | 1 | DS-CC | hgu133a | none | DS-CC 3 |
| 6Twin.CEL | 6 | 2 | DS-twin | hgu133plus2 | none | DS-twin 6 |
| 10358_WBS.CEL | 1 | 8 | WBS | hgu133plus2 | WBS | WBS 1 |
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| E-TABM-263-raw-cel-1515485831.cel | 10 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |