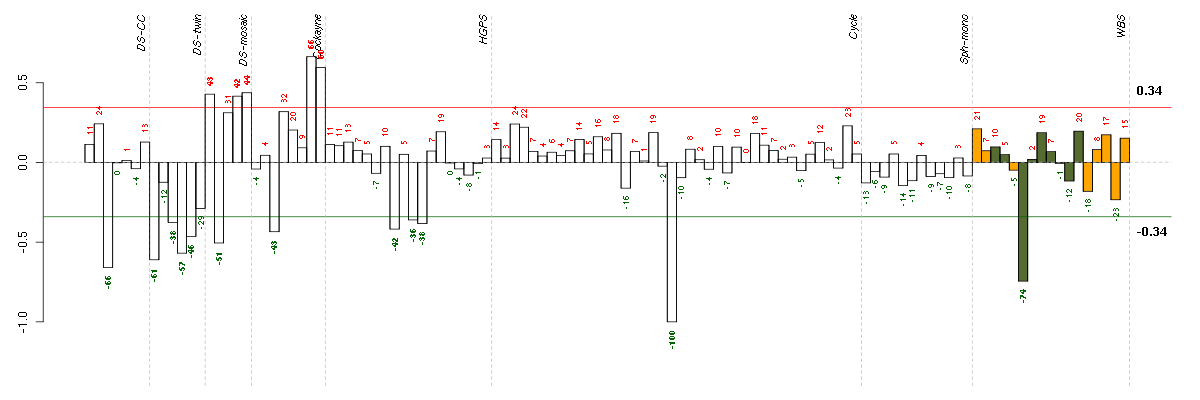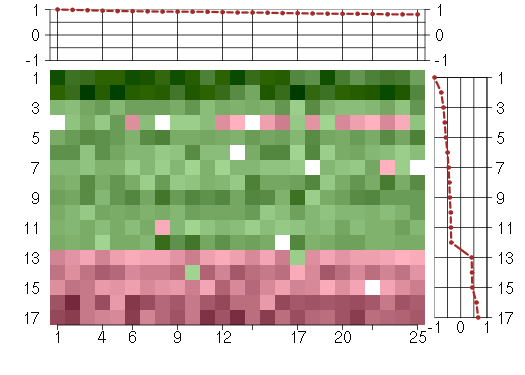

Under-expression is coded with green,
over-expression with red color.



| Id | Pvalue | ExpCount | Count | Size |
|---|---|---|---|---|
| miR-320/320abcd | 3.572e-02 | 1.447 | 9 | 377 |
ATP13A3ATPase type 13A3 (219558_at), score: 0.94 ATP6V1AATPase, H+ transporting, lysosomal 70kDa, V1 subunit A (201971_s_at), score: 0.83 CDV3CDV3 homolog (mouse) (213548_s_at), score: 0.97 COPAcoatomer protein complex, subunit alpha (214336_s_at), score: 1 CPDcarboxypeptidase D (201942_s_at), score: 0.92 DDX3XDEAD (Asp-Glu-Ala-Asp) box polypeptide 3, X-linked (201211_s_at), score: 0.93 DHX9DEAH (Asp-Glu-Ala-His) box polypeptide 9 (212105_s_at), score: 0.99 FN1fibronectin 1 (214701_s_at), score: 0.91 GNSglucosamine (N-acetyl)-6-sulfatase (203676_at), score: 0.91 GTF2Igeneral transcription factor II, i (210892_s_at), score: 0.89 HSPA4heat shock 70kDa protein 4 (211016_x_at), score: 0.81 IL6STinterleukin 6 signal transducer (gp130, oncostatin M receptor) (204864_s_at), score: 0.84 MAT2Amethionine adenosyltransferase II, alpha (200769_s_at), score: 0.81 MAXMYC associated factor X (210734_x_at), score: 0.88 PAFAH1B1platelet-activating factor acetylhydrolase, isoform Ib, alpha subunit 45kDa (211547_s_at), score: 0.83 PCDHGA11protocadherin gamma subfamily A, 11 (211876_x_at), score: 0.95 PCDHGA3protocadherin gamma subfamily A, 3 (216352_x_at), score: 0.86 PHTF2putative homeodomain transcription factor 2 (217097_s_at), score: 0.88 PTBP1polypyrimidine tract binding protein 1 (212016_s_at), score: 0.84 RAB5ARAB5A, member RAS oncogene family (206113_s_at), score: 0.92 SCAMP1secretory carrier membrane protein 1 (206667_s_at), score: 0.85 SCARB2scavenger receptor class B, member 2 (201647_s_at), score: 0.83 STIP1stress-induced-phosphoprotein 1 (212009_s_at), score: 0.81 TGFBR2transforming growth factor, beta receptor II (70/80kDa) (207334_s_at), score: 0.87 WACWW domain containing adaptor with coiled-coil (219679_s_at), score: 0.93
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515486031.cel | 20 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 5042_CNTL.CEL | 6 | 8 | WBS | hgu133plus2 | none | WBS 1 |
| ctrl c 08-03.CEL | 3 | 1 | DS-CC | hgu133a | none | DS-CC 3 |
| 1Twin.CEL | 1 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 1 |
| 4Twin.CEL | 4 | 2 | DS-twin | hgu133plus2 | none | DS-twin 4 |
| 46B.CEL | 2 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 2 |
| 5CTwin.CEL | 5 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 5 |
| E-GEOD-3407-raw-cel-1437949655.cel | 3 | 4 | Cockayne | hgu133a | none | CSB |
| E-GEOD-3860-raw-cel-1561690304.cel | 8 | 5 | HGPS | hgu133a | none | GMO8398C |
| E-GEOD-3860-raw-cel-1561690352.cel | 11 | 5 | HGPS | hgu133a | HGPS | AG11498 |
| 3Twin.CEL | 3 | 2 | DS-twin | hgu133plus2 | Down | DS-twin 3 |
| E-GEOD-3860-raw-cel-1561690344.cel | 10 | 5 | HGPS | hgu133a | none | GM00038C |
| 47B.CEL | 4 | 3 | DS-mosaic | hgu133plus2 | Down mosaic | DS-mosaic 4 |
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| 47C.CEL | 5 | 3 | DS-mosaic | hgu133plus2 | Down mosaic | DS-mosaic 5 |
| E-GEOD-3407-raw-cel-1437949938.cel | 8 | 4 | Cockayne | hgu133a | none | CSB |
| E-GEOD-3407-raw-cel-1437949854.cel | 7 | 4 | Cockayne | hgu133a | CS | eGFP |