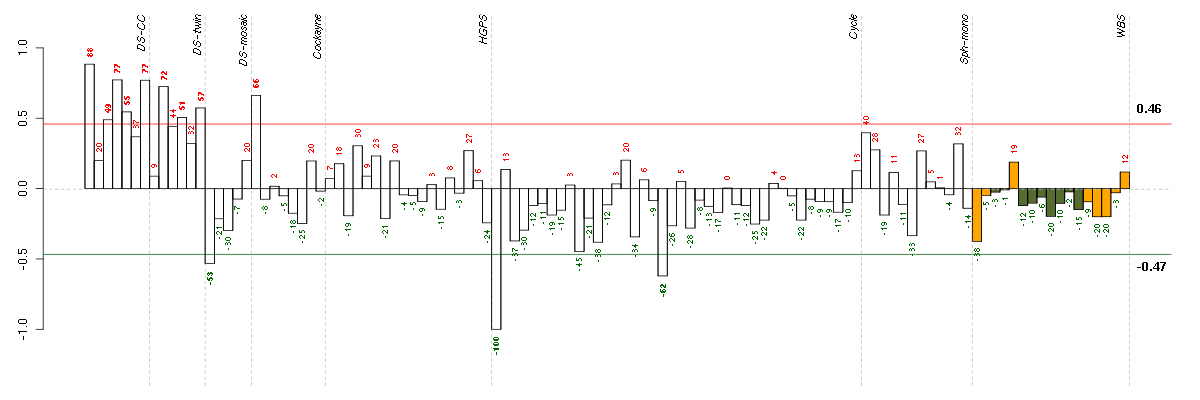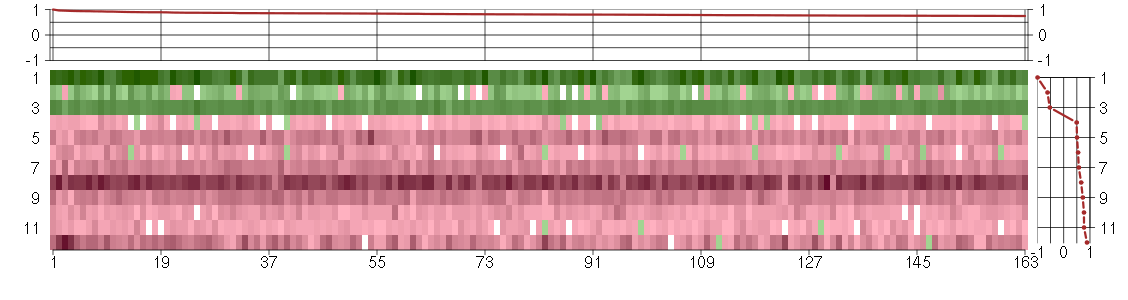

Under-expression is coded with green,
over-expression with red color.

system process
A multicellular organismal process carried out by any of the organs or tissues in an organ system. An organ system is a regularly interacting or interdependent group of organs or tissues that work together to carry out a biological objective.
signal transduction
The cascade of processes by which a signal interacts with a receptor, causing a change in the level or activity of a second messenger or other downstream target, and ultimately effecting a change in the functioning of the cell.
cell communication
Any process that mediates interactions between a cell and its surroundings. Encompasses interactions such as signaling or attachment between one cell and another cell, between a cell and an extracellular matrix, or between a cell and any other aspect of its environment.
cell surface receptor linked signal transduction
Any series of molecular signals initiated by the binding of an extracellular ligand to a receptor on the surface of the target cell.
G-protein coupled receptor protein signaling pathway
The series of molecular signals generated as a consequence of a G-protein coupled receptor binding to its physiological ligand.
neurological system process
A organ system process carried out by any of the organs or tissues of neurological system.
digestion
The whole of the physical, chemical, and biochemical processes carried out by multicellular organisms to break down ingested nutrients into components that may be easily absorbed and directed into metabolism.
sensory perception
The series of events required for an organism to receive a sensory stimulus, convert it to a molecular signal, and recognize and characterize the signal.
sensory perception of chemical stimulus
The series of events required for an organism to receive a sensory chemical stimulus, convert it to a molecular signal, and recognize and characterize the signal.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
cellular process
Any process that is carried out at the cellular level, but not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
multicellular organismal process
Any biological process, occurring at the level of a multicellular organism, pertinent to its function.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
cognition
The operation of the mind by which an organism becomes aware of objects of thought or perception; it includes the mental activities associated with thinking, learning, and memory.
biological regulation
Any process that modulates the frequency, rate or extent of any biological process, quality or function.
all
This term is the most general term possible
regulation of cellular process
Any process that modulates the frequency, rate or extent of a cellular process, any of those that are carried out at the cellular level, but are not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
regulation of biological process
Any process that modulates the frequency, rate or extent of a biological process. Biological processes are regulated by many means; examples include the control of gene expression, protein modification or interaction with a protein or substrate molecule.
signal transduction
The cascade of processes by which a signal interacts with a receptor, causing a change in the level or activity of a second messenger or other downstream target, and ultimately effecting a change in the functioning of the cell.

plasma membrane
The membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
membrane
Double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
integral to membrane
Penetrating at least one phospholipid bilayer of a membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer. When used to describe a protein, indicates that all or part of the peptide sequence is embedded in the membrane.
extracellular region
The space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers the host cell environment outside an intracellular parasite.
integral to plasma membrane
Penetrating at least one phospholipid bilayer of a plasma membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
cell
The basic structural and functional unit of all organisms. Includes the plasma membrane and any external encapsulating structures such as the cell wall and cell envelope.
intrinsic to membrane
Located in a membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
intrinsic to plasma membrane
Located in the plasma membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
plasma membrane part
Any constituent part of the plasma membrane, the membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
all
This term is the most general term possible
cell part
Any constituent part of a cell, the basic structural and functional unit of all organisms.
membrane part
Any constituent part of a membrane, a double layer of lipid molecules that encloses all cells, and, in eukaryotes, many organelles; may be a single or double lipid bilayer; also includes associated proteins.
plasma membrane part
Any constituent part of the plasma membrane, the membrane surrounding a cell that separates the cell from its external environment. It consists of a phospholipid bilayer and associated proteins.
intrinsic to plasma membrane
Located in the plasma membrane such that some covalently attached portion of the gene product, for example part of a peptide sequence or some other covalently attached moiety such as a GPI anchor, spans or is embedded in one or both leaflets of the membrane.
integral to plasma membrane
Penetrating at least one phospholipid bilayer of a plasma membrane. May also refer to the state of being buried in the bilayer with no exposure outside the bilayer.

molecular_function
Elemental activities, such as catalysis or binding, describing the actions of a gene product at the molecular level. A given gene product may exhibit one or more molecular functions.
catalytic activity
Catalysis of a biochemical reaction at physiological temperatures. In biologically catalyzed reactions, the reactants are known as substrates, and the catalysts are naturally occurring macromolecular substances known as enzymes. Enzymes possess specific binding sites for substrates, and are usually composed wholly or largely of protein, but RNA that has catalytic activity (ribozyme) is often also regarded as enzymatic.
monooxygenase activity
Catalysis of the incorporation of one atom from molecular oxygen into a compound and the reduction of the other atom of oxygen to water.
signal transducer activity
Mediates the transfer of a signal from the outside to the inside of a cell by means other than the introduction of the signal molecule itself into the cell.
receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity.
transmembrane receptor activity
Combining with an extracellular or intracellular messenger to initiate a change in cell activity, and spanning to the membrane of either the cell or an organelle.
G-protein coupled receptor activity
A receptor that binds an extracellular ligand and transmits the signal to a heterotrimeric G-protein complex. These receptors are characteristically seven-transmembrane receptors and are made up of hetero- or homodimers.
transporter activity
Enables the directed movement of substances (such as macromolecules, small molecules, ions) into, out of, within or between cells.
transmembrane transporter activity
Catalysis of the transfer of a substance from one side of a membrane to the other.
organic acid:sodium symporter activity
Catalysis of the transfer of a solute or solutes from one side of a membrane to the other according to the reaction: organic acid(out) + Na+(out) = organic acid(in) + Na+(in).
organic acid transmembrane transporter activity
Catalysis of the transfer of organic acids, any acidic compound containing carbon in covalent linkage, from one side of the membrane to the other.
secondary active transmembrane transporter activity
Catalysis of the transfer of a solute from one side of a membrane to the other, up its concentration gradient. The transporter binds the solute and undergoes a series of conformational changes. Transport works equally well in either direction and is driven by a chemiosmotic source of energy. Chemiosmotic sources of energy include uniport, symport or antiport.
oxidoreductase activity
Catalysis of an oxidation-reduction (redox) reaction, a reversible chemical reaction in which the oxidation state of an atom or atoms within a molecule is altered. One substrate acts as a hydrogen or electron donor and becomes oxidized, while the other acts as hydrogen or electron acceptor and becomes reduced.
cation transmembrane transporter activity
Catalysis of the transfer of cation from one side of the membrane to the other.
ion transmembrane transporter activity
Catalysis of the transfer of an ion from one side of a membrane to the other.
symporter activity
Enables the active transport of a solute across a membrane by a mechanism whereby two or more species are transported together in the same direction in a tightly coupled process not directly linked to a form of energy other than chemiosmotic energy.
solute:cation symporter activity
Catalysis of the transfer of a solute or solutes from one side of a membrane to the other according to the reaction: solute(out) + cation(out) = solute(in) + cation(in).
solute:sodium symporter activity
Catalysis of the transfer of a solute or solutes from one side of a membrane to the other according to the reaction: solute(out) + Na+(out) = solute(in) + Na+(in).
active transmembrane transporter activity
Catalysis of the transfer of a specific substance or related group of substances from one side of a membrane to the other, up the solute's concentration gradient. The transporter binds the solute and undergoes a series of conformational changes. Transport works equally well in either direction.
substrate-specific transmembrane transporter activity
Catalysis of the transfer of a specific substance or group of related substances from one side of a membrane to the other.
substrate-specific transporter activity
Enables the directed movement of a specific substance or group of related substances (such as macromolecules, small molecules, ions) into, out of, within or between cells.
molecular transducer activity
The molecular function that accepts an input of one form and creates an output of a different form.
all
This term is the most general term possible
substrate-specific transmembrane transporter activity
Catalysis of the transfer of a specific substance or group of related substances from one side of a membrane to the other.
solute:cation symporter activity
Catalysis of the transfer of a solute or solutes from one side of a membrane to the other according to the reaction: solute(out) + cation(out) = solute(in) + cation(in).
organic acid:sodium symporter activity
Catalysis of the transfer of a solute or solutes from one side of a membrane to the other according to the reaction: organic acid(out) + Na+(out) = organic acid(in) + Na+(in).
| Id | Pvalue | ExpCount | Count | Size | Term |
|---|---|---|---|---|---|
| 04080 | 4.998e-02 | 1.614 | 7 | 83 | Neuroactive ligand-receptor interaction |
A1CFAPOBEC1 complementation factor (220951_s_at), score: 0.8 ABCG4ATP-binding cassette, sub-family G (WHITE), member 4 (207593_at), score: 0.8 ACSL6acyl-CoA synthetase long-chain family member 6 (211207_s_at), score: 0.79 ADAMTS9ADAM metallopeptidase with thrombospondin type 1 motif, 9 (220287_at), score: 0.83 ADAMTSL4ADAMTS-like 4 (220578_at), score: 0.75 ADRBK2adrenergic, beta, receptor kinase 2 (204184_s_at), score: 0.78 AGMATagmatine ureohydrolase (agmatinase) (219792_at), score: 0.85 AIPL1aryl hydrocarbon receptor interacting protein-like 1 (219977_at), score: 0.81 ALDOBaldolase B, fructose-bisphosphate (214423_x_at), score: 0.82 APOMapolipoprotein M (205682_x_at), score: 0.87 ATRNL1attractin-like 1 (213744_at), score: 0.78 BMP10bone morphogenetic protein 10 (208292_at), score: 0.78 BRS3bombesin-like receptor 3 (207369_at), score: 0.81 BTNL3butyrophilin-like 3 (217207_s_at), score: 0.89 C10orf59chromosome 10 open reading frame 59 (220564_at), score: 0.76 C1orf61chromosome 1 open reading frame 61 (205103_at), score: 0.86 C2orf54chromosome 2 open reading frame 54 (220149_at), score: 0.83 C6orf134chromosome 6 open reading frame 134 (218874_s_at), score: 0.75 C7orf28Achromosome 7 open reading frame 28A (201974_s_at), score: 0.86 C8orf17chromosome 8 open reading frame 17 (208266_at), score: 0.87 C8orf60chromosome 8 open reading frame 60 (220712_at), score: 0.82 CA4carbonic anhydrase IV (206209_s_at), score: 0.87 CAPN9calpain 9 (210641_at), score: 0.76 CASZ1castor zinc finger 1 (220015_at), score: 0.84 CATSPER2cation channel, sperm associated 2 (217588_at), score: 0.81 CCDC9coiled-coil domain containing 9 (206257_at), score: 0.82 CD53CD53 molecule (203416_at), score: 0.85 CEACAM1carcinoembryonic antigen-related cell adhesion molecule 1 (biliary glycoprotein) (211883_x_at), score: 0.77 CEACAM5carcinoembryonic antigen-related cell adhesion molecule 5 (201884_at), score: 0.85 CEACAM6carcinoembryonic antigen-related cell adhesion molecule 6 (non-specific cross reacting antigen) (211657_at), score: 0.9 CHD2chromodomain helicase DNA binding protein 2 (203461_at), score: 0.82 CHN2chimerin (chimaerin) 2 (207486_x_at), score: 0.76 CHST4carbohydrate (N-acetylglucosamine 6-O) sulfotransferase 4 (220446_s_at), score: 0.83 CLDN10claudin 10 (205328_at), score: 0.92 CLDN18claudin 18 (214135_at), score: 0.84 CLEC4MC-type lectin domain family 4, member M (207995_s_at), score: 0.92 CNKSR2connector enhancer of kinase suppressor of Ras 2 (206731_at), score: 0.76 COL11A2collagen, type XI, alpha 2 (216993_s_at), score: 0.82 CSN2casein beta (207951_at), score: 0.84 CTSEcathepsin E (205927_s_at), score: 0.78 CYBBcytochrome b-245, beta polypeptide (203923_s_at), score: 0.9 CYP2A6cytochrome P450, family 2, subfamily A, polypeptide 6 (211295_x_at), score: 0.81 CYP2C9cytochrome P450, family 2, subfamily C, polypeptide 9 (214421_x_at), score: 0.8 CYP2E1cytochrome P450, family 2, subfamily E, polypeptide 1 (1431_at), score: 0.83 DAOD-amino-acid oxidase (206878_at), score: 0.8 DHRS9dehydrogenase/reductase (SDR family) member 9 (219799_s_at), score: 0.93 DNASE1L3deoxyribonuclease I-like 3 (205554_s_at), score: 0.84 DOHHdeoxyhypusine hydroxylase/monooxygenase (208141_s_at), score: 0.78 DRD5dopamine receptor D5 (208486_at), score: 0.85 EDAectodysplasin A (211130_x_at), score: 0.76 EDARectodysplasin A receptor (220048_at), score: 0.81 FAM38Bfamily with sequence similarity 38, member B (219602_s_at), score: 0.84 FAM75A3family with sequence similarity 75, member A3 (215935_at), score: 0.77 FGL1fibrinogen-like 1 (205305_at), score: 0.75 FMO4flavin containing monooxygenase 4 (206263_at), score: 0.8 GABRR2gamma-aminobutyric acid (GABA) receptor, rho 2 (208217_at), score: 0.81 GLTSCR1glioma tumor suppressor candidate region gene 1 (219445_at), score: 0.76 GNG4guanine nucleotide binding protein (G protein), gamma 4 (205184_at), score: 0.76 GNRHRgonadotropin-releasing hormone receptor (216341_s_at), score: 0.82 GP1BAglycoprotein Ib (platelet), alpha polypeptide (207389_at), score: 0.76 GP6glycoprotein VI (platelet) (220336_s_at), score: 0.75 GPATCH4G patch domain containing 4 (220596_at), score: 0.76 GPR12G protein-coupled receptor 12 (214558_at), score: 0.89 GPR157G protein-coupled receptor 157 (220901_at), score: 0.85 GRHL2grainyhead-like 2 (Drosophila) (219388_at), score: 0.75 GRM6glutamate receptor, metabotropic 6 (208035_at), score: 0.79 HABP2hyaluronan binding protein 2 (206010_at), score: 0.77 HAO2hydroxyacid oxidase 2 (long chain) (220801_s_at), score: 0.75 HAVCR1hepatitis A virus cellular receptor 1 (207052_at), score: 0.74 HIST1H4Dhistone cluster 1, H4d (208076_at), score: 0.8 HOXD4homeobox D4 (205522_at), score: 0.81 HS3ST2heparan sulfate (glucosamine) 3-O-sulfotransferase 2 (219697_at), score: 0.76 INE1inactivation escape 1 (non-protein coding) (207252_at), score: 0.78 ITIH2inter-alpha (globulin) inhibitor H2 (204987_at), score: 0.85 ITKIL2-inducible T-cell kinase (211339_s_at), score: 0.76 KCNIP1Kv channel interacting protein 1 (221307_at), score: 0.75 KCNJ16potassium inwardly-rectifying channel, subfamily J, member 16 (219564_at), score: 0.86 KCNMB4potassium large conductance calcium-activated channel, subfamily M, beta member 4 (219287_at), score: 0.8 KIAA0748KIAA0748 (219724_s_at), score: 0.81 KIF5Akinesin family member 5A (205318_at), score: 0.8 KIR3DX1killer cell immunoglobulin-like receptor, three domains, X1 (216428_x_at), score: 0.89 KLF1Kruppel-like factor 1 (erythroid) (210504_at), score: 0.93 KLHL4kelch-like 4 (Drosophila) (214591_at), score: 0.76 KMOkynurenine 3-monooxygenase (kynurenine 3-hydroxylase) (205307_s_at), score: 0.79 KNG1kininogen 1 (217512_at), score: 0.75 KRT2keratin 2 (207908_at), score: 0.89 LHX3LIM homeobox 3 (221670_s_at), score: 0.9 LOC100188945cell division cycle associated 4 pseudogene (215109_at), score: 0.85 LOC389906similar to Serine/threonine-protein kinase PRKX (Protein kinase PKX1) (59433_at), score: 0.81 LOC643293FLJ44451 pseudogene (215057_at), score: 0.86 LONRF3LON peptidase N-terminal domain and ring finger 3 (220009_at), score: 0.85 MALmal, T-cell differentiation protein (204777_s_at), score: 0.76 MAPK8IP2mitogen-activated protein kinase 8 interacting protein 2 (205050_s_at), score: 0.77 MBmyoglobin (204179_at), score: 0.91 MEP1Bmeprin A, beta (207251_at), score: 0.75 MFNGMFNG O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase (204153_s_at), score: 0.78 MMP9matrix metallopeptidase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) (203936_s_at), score: 0.84 MMRN2multimerin 2 (219091_s_at), score: 0.96 MOBPmyelin-associated oligodendrocyte basic protein (210193_at), score: 0.83 MSTP9macrophage stimulating, pseudogene 9 (213382_at), score: 0.87 MUC3Amucin 3A, cell surface associated (217117_x_at), score: 0.8 MYL4myosin, light chain 4, alkali; atrial, embryonic (210088_x_at), score: 0.77 MYO1Amyosin IA (211916_s_at), score: 0.81 N4BP3Nedd4 binding protein 3 (214775_at), score: 0.89 NCRNA00093non-protein coding RNA 93 (210723_x_at), score: 0.76 NEUROD2neurogenic differentiation 2 (210271_at), score: 0.75 NINninein (GSK3B interacting protein) (219285_s_at), score: 0.77 NOTCH4Notch homolog 4 (Drosophila) (205247_at), score: 0.77 NRN1neuritin 1 (218625_at), score: 0.87 NTNG1netrin G1 (206713_at), score: 0.84 NTRK2neurotrophic tyrosine kinase, receptor, type 2 (207152_at), score: 0.79 OCLMoculomedin (208274_at), score: 0.8 OR10C1olfactory receptor, family 10, subfamily C, member 1 (221339_at), score: 0.79 OR1D2olfactory receptor, family 1, subfamily D, member 2 (221464_at), score: 0.76 OR7E24olfactory receptor, family 7, subfamily E, member 24 (215463_at), score: 0.94 OVOL2ovo-like 2 (Drosophila) (211778_s_at), score: 0.78 PECAM1platelet/endothelial cell adhesion molecule (208982_at), score: 1 PEG3paternally expressed 3 (209242_at), score: 0.87 PGCprogastricsin (pepsinogen C) (205261_at), score: 0.81 PGK2phosphoglycerate kinase 2 (217009_at), score: 0.75 PIPOXpipecolic acid oxidase (221605_s_at), score: 0.78 PLUNCpalate, lung and nasal epithelium associated (220542_s_at), score: 0.86 PNPLA2patatin-like phospholipase domain containing 2 (212705_x_at), score: 0.77 PON1paraoxonase 1 (206344_at), score: 0.76 POU4F1POU class 4 homeobox 1 (206940_s_at), score: 0.8 PRB3proline-rich protein BstNI subfamily 3 (206998_x_at), score: 0.79 PROL1proline rich, lacrimal 1 (208004_at), score: 0.78 PROX1prospero homeobox 1 (207401_at), score: 0.82 PRSS7protease, serine, 7 (enterokinase) (217269_s_at), score: 0.94 PYHIN1pyrin and HIN domain family, member 1 (216748_at), score: 0.88 RAB26RAB26, member RAS oncogene family (50965_at), score: 0.76 RETret proto-oncogene (205879_x_at), score: 0.86 RNASE2ribonuclease, RNase A family, 2 (liver, eosinophil-derived neurotoxin) (206111_at), score: 0.75 ROS1c-ros oncogene 1 , receptor tyrosine kinase (207569_at), score: 0.85 RP11-35N6.1plasticity related gene 3 (219732_at), score: 0.93 RPGRIP1retinitis pigmentosa GTPase regulator interacting protein 1 (206608_s_at), score: 0.93 RPH3Arabphilin 3A homolog (mouse) (205230_at), score: 0.79 RRHretinal pigment epithelium-derived rhodopsin homolog (208314_at), score: 0.85 S100A14S100 calcium binding protein A14 (218677_at), score: 0.87 SCN10Asodium channel, voltage-gated, type X, alpha subunit (208578_at), score: 0.82 SEMA6Dsema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D (220574_at), score: 0.78 SERPINA1serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 1 (211429_s_at), score: 0.82 SERPINB3serpin peptidase inhibitor, clade B (ovalbumin), member 3 (209719_x_at), score: 0.77 SLC10A1solute carrier family 10 (sodium/bile acid cotransporter family), member 1 (207185_at), score: 0.87 SLC10A2solute carrier family 10 (sodium/bile acid cotransporter family), member 2 (207095_at), score: 0.8 SLC15A1solute carrier family 15 (oligopeptide transporter), member 1 (211349_at), score: 0.85 SLC1A2solute carrier family 1 (glial high affinity glutamate transporter), member 2 (208389_s_at), score: 0.77 SLC1A7solute carrier family 1 (glutamate transporter), member 7 (210923_at), score: 0.81 SLC26A10solute carrier family 26, member 10 (214951_at), score: 0.8 SP140SP140 nuclear body protein (207777_s_at), score: 0.77 STXBP5Lsyntaxin binding protein 5-like (215518_at), score: 0.84 TAAR2trace amine associated receptor 2 (221394_at), score: 0.75 TACR1tachykinin receptor 1 (208048_at), score: 0.88 TAS2R16taste receptor, type 2, member 16 (221444_at), score: 0.79 TAS2R9taste receptor, type 2, member 9 (221461_at), score: 0.79 TBX21T-box 21 (220684_at), score: 0.85 TBX6T-box 6 (207684_at), score: 0.87 TP63tumor protein p63 (209863_s_at), score: 0.84 TRIM10tripartite motif-containing 10 (56748_at), score: 0.8 TRIM45tripartite motif-containing 45 (219923_at), score: 0.75 TRIM49tripartite motif-containing 49 (221154_at), score: 0.75 VGFVGF nerve growth factor inducible (205586_x_at), score: 0.95 VPRBPVpr (HIV-1) binding protein (204377_s_at), score: 0.75
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515485651.cel | 1 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| 46A.CEL | 1 | 3 | DS-mosaic | hgu133plus2 | none | DS-mosaic 1 |
| ctrl c 08-03.CEL | 3 | 1 | DS-CC | hgu133a | none | DS-CC 3 |
| 4Twin.CEL | 4 | 2 | DS-twin | hgu133plus2 | none | DS-twin 4 |
| t21b 08-03.CEL | 5 | 1 | DS-CC | hgu133a | Down | DS-CC 5 |
| 6Twin.CEL | 6 | 2 | DS-twin | hgu133plus2 | none | DS-twin 6 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| 2Twin.CEL | 2 | 2 | DS-twin | hgu133plus2 | none | DS-twin 2 |
| t21d 08-03.CEL | 7 | 1 | DS-CC | hgu133a | Down | DS-CC 7 |
| t21a 08-03.CEL | 4 | 1 | DS-CC | hgu133a | Down | DS-CC 4 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |