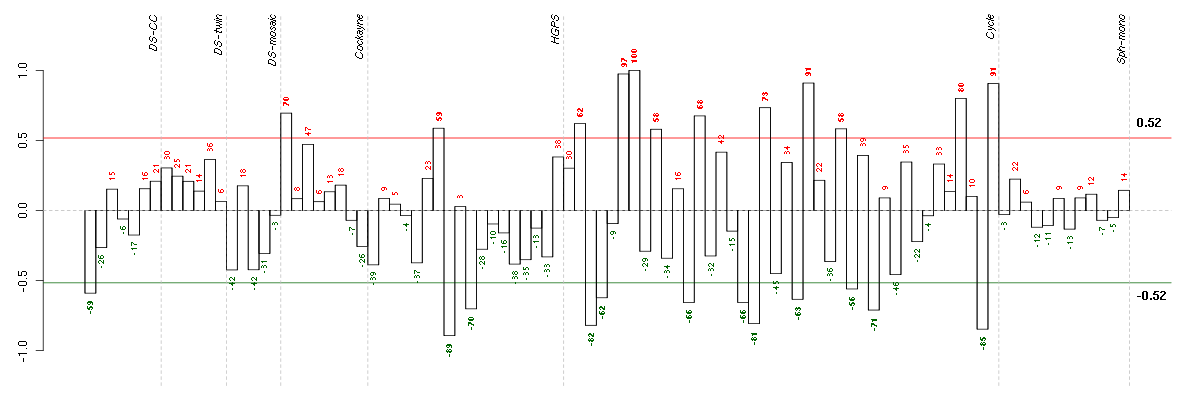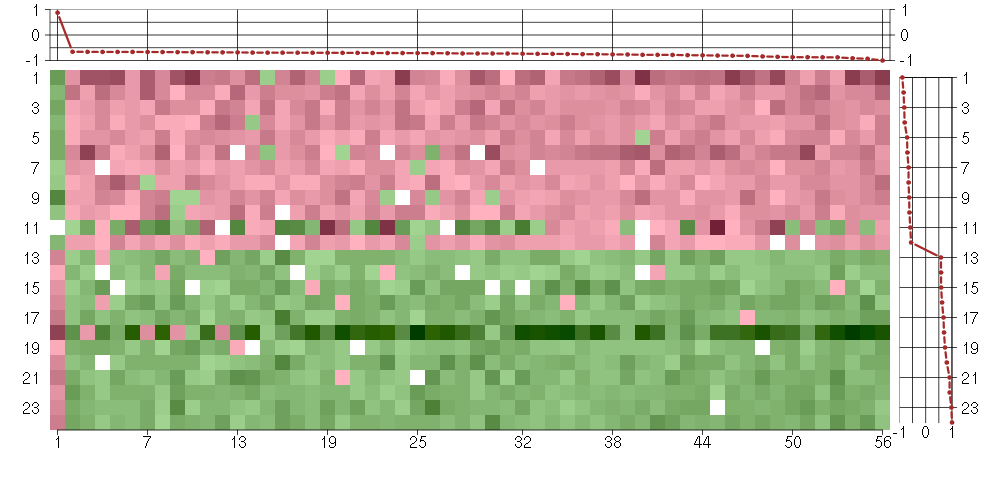

Under-expression is coded with green,
over-expression with red color.



ABCD1ATP-binding cassette, sub-family D (ALD), member 1 (205142_x_at), score: -0.73 ADCK2aarF domain containing kinase 2 (221893_s_at), score: -0.67 ARSAarylsulfatase A (204443_at), score: -0.75 ATN1atrophin 1 (40489_at), score: -0.86 B3GAT3beta-1,3-glucuronyltransferase 3 (glucuronosyltransferase I) (203452_at), score: -0.71 BAT2HLA-B associated transcript 2 (212081_x_at), score: -0.71 BTBD2BTB (POZ) domain containing 2 (207722_s_at), score: -0.8 C16orf57chromosome 16 open reading frame 57 (218060_s_at), score: -0.66 CDC42BPBCDC42 binding protein kinase beta (DMPK-like) (217849_s_at), score: -0.87 CDC42EP1CDC42 effector protein (Rho GTPase binding) 1 (204693_at), score: -0.74 CICcapicua homolog (Drosophila) (212784_at), score: -0.78 CIZ1CDKN1A interacting zinc finger protein 1 (205516_x_at), score: -0.69 CNOT3CCR4-NOT transcription complex, subunit 3 (203239_s_at), score: -0.76 CRTC3CREB regulated transcription coactivator 3 (218648_at), score: -0.69 DMWDdystrophia myotonica, WD repeat containing (33768_at), score: -0.67 FHOD1formin homology 2 domain containing 1 (218530_at), score: -0.7 FLJ12529pre-mRNA cleavage factor I, 59 kDa subunit (217866_at), score: -0.73 FOXK2forkhead box K2 (203064_s_at), score: -0.88 FURINfurin (paired basic amino acid cleaving enzyme) (201945_at), score: -0.66 GRINAglutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1 (glutamate binding) (212090_at), score: -0.75 HMGA1high mobility group AT-hook 1 (210457_x_at), score: -0.73 IDUAiduronidase, alpha-L- (205059_s_at), score: -0.82 JUNDjun D proto-oncogene (203751_x_at), score: -0.71 LEPREL2leprecan-like 2 (204854_at), score: -0.7 MAP1Smicrotubule-associated protein 1S (218522_s_at), score: -1 MAP2K3mitogen-activated protein kinase kinase 3 (207667_s_at), score: -0.7 MAP3K6mitogen-activated protein kinase kinase kinase 6 (219278_at), score: -0.67 MAP7D1MAP7 domain containing 1 (217943_s_at), score: -0.76 MAST2microtubule associated serine/threonine kinase 2 (211593_s_at), score: -0.67 NBL1neuroblastoma, suppression of tumorigenicity 1 (37005_at), score: -0.66 NFATC4nuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 4 (205897_at), score: -0.68 PARVBparvin, beta (37965_at), score: -0.72 PCDHG@protocadherin gamma cluster (215836_s_at), score: -0.75 PCDHGA1protocadherin gamma subfamily A, 1 (209079_x_at), score: -0.81 PIP5K1Cphosphatidylinositol-4-phosphate 5-kinase, type I, gamma (212518_at), score: -0.91 POLR2Apolymerase (RNA) II (DNA directed) polypeptide A, 220kDa (202725_at), score: -0.74 POM121POM121 membrane glycoprotein (rat) (212178_s_at), score: -0.78 PTPROprotein tyrosine phosphatase, receptor type, O (211600_at), score: -0.72 PXNpaxillin (211823_s_at), score: -0.69 RAD54L2RAD54-like 2 (S. cerevisiae) (213205_s_at), score: -0.66 RBM15BRNA binding motif protein 15B (202689_at), score: -0.7 RPL18AP6ribosomal protein L18a pseudogene 6 (216383_at), score: 0.87 SCAMP4secretory carrier membrane protein 4 (213244_at), score: -0.93 SF1splicing factor 1 (208313_s_at), score: -0.79 SHBSrc homology 2 domain containing adaptor protein B (204657_s_at), score: -0.66 SOLHsmall optic lobes homolog (Drosophila) (204275_at), score: -0.87 SPRY4sprouty homolog 4 (Drosophila) (221489_s_at), score: -0.68 SSBP3single stranded DNA binding protein 3 (217991_x_at), score: -0.71 STRN4striatin, calmodulin binding protein 4 (217903_at), score: -0.78 TAPBPTAP binding protein (tapasin) (208829_at), score: -0.73 TSKUtsukushin (218245_at), score: -0.71 UBTD1ubiquitin domain containing 1 (219172_at), score: -0.68 VEGFBvascular endothelial growth factor B (203683_s_at), score: -0.88 VPS37Bvacuolar protein sorting 37 homolog B (S. cerevisiae) (221704_s_at), score: -0.84 XAB2XPA binding protein 2 (218110_at), score: -0.69 ZFP36L1zinc finger protein 36, C3H type-like 1 (211965_at), score: -0.81
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-GEOD-3860-raw-cel-1561690304.cel | 8 | 5 | HGPS | hgu133a | none | GMO8398C |
| E-TABM-263-raw-cel-1515486411.cel | 39 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485691.cel | 3 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485991.cel | 18 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486211.cel | 29 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3860-raw-cel-1561690344.cel | 10 | 5 | HGPS | hgu133a | none | GM00038C |
| E-TABM-263-raw-cel-1515485871.cel | 12 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485971.cel | 17 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486071.cel | 22 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485711.cel | 4 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| ctrl a 08-03.CEL | 1 | 1 | DS-CC | hgu133a | none | DS-CC 1 |
| E-TABM-263-raw-cel-1515486171.cel | 27 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485811.cel | 9 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486151.cel | 26 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3860-raw-cel-1561690272.cel | 7 | 5 | HGPS | hgu133a | HGPS | AG11498 |
| E-TABM-263-raw-cel-1515485671.cel | 2 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485891.cel | 13 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-GEOD-3407-raw-cel-1437949557.cel | 1 | 4 | Cockayne | hgu133a | CS | eGFP |
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486371.cel | 37 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486431.cel | 40 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486091.cel | 23 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485751.cel | 6 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515485771.cel | 7 | 6 | Cycle | hgu133a2 | none | Cycle 1 |