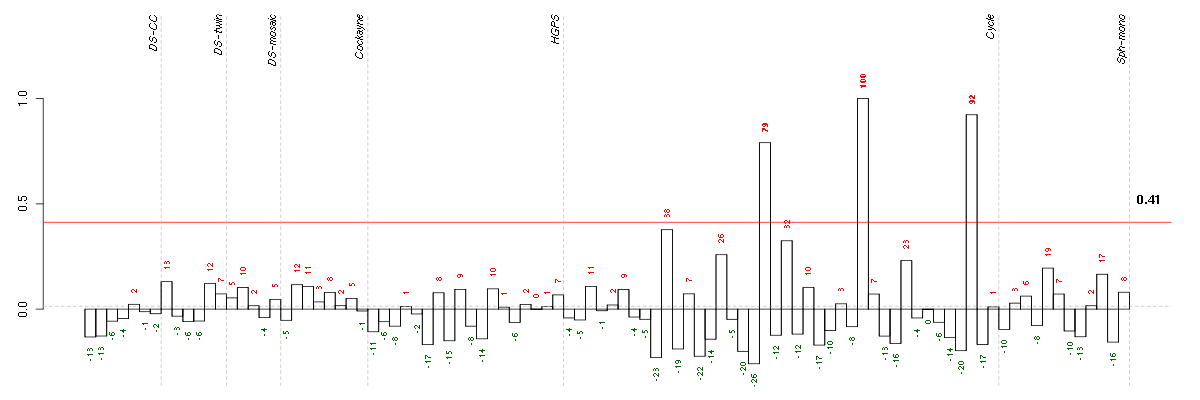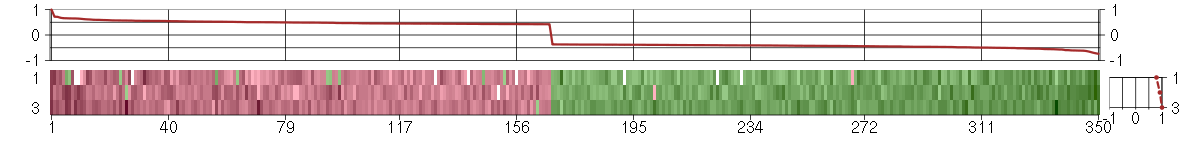

Under-expression is coded with green,
over-expression with red color.

cell recognition
The process by which a cell in a multicellular organism interprets its surroundings.
biological_process
Any process specifically pertinent to the functioning of integrated living units: cells, tissues, organs, and organisms. A process is a collection of molecular events with a defined beginning and end.
cellular process
Any process that is carried out at the cellular level, but not necessarily restricted to a single cell. For example, cell communication occurs among more than one cell, but occurs at the cellular level.
cell-cell recognition
Cell recognition between cells, usually involving the formation of specialized cell junctions.
all
This term is the most general term possible

extracellular region
The space external to the outermost structure of a cell. For cells without external protective or external encapsulating structures this refers to space outside of the plasma membrane. This term covers the host cell environment outside an intracellular parasite.
cellular_component
The part of a cell or its extracellular environment in which a gene product is located. A gene product may be located in one or more parts of a cell and its location may be as specific as a particular macromolecular complex, that is, a stable, persistent association of macromolecules that function together.
all
This term is the most general term possible

ABCA2ATP-binding cassette, sub-family A (ABC1), member 2 (212772_s_at), score: -0.42 ACANaggrecan (205679_x_at), score: 0.53 ADAM21ADAM metallopeptidase domain 21 (207665_at), score: 0.42 ADAMTS6ADAM metallopeptidase with thrombospondin type 1 motif, 6 (220866_at), score: -0.54 ADCK2aarF domain containing kinase 2 (221893_s_at), score: -0.39 ADSSadenylosuccinate synthase (221761_at), score: -0.39 AFG3L2AFG3 ATPase family gene 3-like 2 (yeast) (202486_at), score: 0.47 AGBL5ATP/GTP binding protein-like 5 (218480_at), score: 0.49 AHI1Abelson helper integration site 1 (221569_at), score: -0.41 ANKRD28ankyrin repeat domain 28 (213035_at), score: 0.44 ANXA10annexin A10 (210143_at), score: -0.57 AOF2amine oxidase (flavin containing) domain 2 (212348_s_at), score: 0.52 APAF1apoptotic peptidase activating factor 1 (204859_s_at), score: -0.45 APOBEC3Fapolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3F (214995_s_at), score: 0.49 ARL17ADP-ribosylation factor-like 17 (210435_at), score: -0.56 ARMC9armadillo repeat containing 9 (219637_at), score: 0.49 ARNT2aryl-hydrocarbon receptor nuclear translocator 2 (202986_at), score: 0.49 ASPNasporin (219087_at), score: -0.43 ASTN2astrotactin 2 (209693_at), score: 0.45 B3GNT2UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2 (219326_s_at), score: -0.37 B3GNTL1UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase-like 1 (213589_s_at), score: 0.72 B4GALT1UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 1 (201883_s_at), score: 0.48 BAALCbrain and acute leukemia, cytoplasmic (218899_s_at), score: -0.43 BAXBCL2-associated X protein (208478_s_at), score: 0.57 BCL2B-cell CLL/lymphoma 2 (203685_at), score: -0.39 BCS1LBCS1-like (yeast) (207618_s_at), score: 0.45 BTBD3BTB (POZ) domain containing 3 (202946_s_at), score: -0.37 BTN3A2butyrophilin, subfamily 3, member A2 (209846_s_at), score: 0.45 C11orf67chromosome 11 open reading frame 67 (221600_s_at), score: 0.49 C12orf52chromosome 12 open reading frame 52 (221777_at), score: 0.56 C14orf45chromosome 14 open reading frame 45 (220173_at), score: -0.38 C17orf39chromosome 17 open reading frame 39 (220058_at), score: 0.59 C1orf9chromosome 1 open reading frame 9 (203429_s_at), score: -0.49 C2CD2LC2CD2-like (204757_s_at), score: -0.49 C2orf49chromosome 2 open reading frame 49 (219662_at), score: 0.44 C6orf62chromosome 6 open reading frame 62 (213875_x_at), score: 0.64 C7orf58chromosome 7 open reading frame 58 (220032_at), score: -0.37 C8orf17chromosome 8 open reading frame 17 (208266_at), score: -0.42 C8orf41chromosome 8 open reading frame 41 (219124_at), score: 0.5 C9orf114chromosome 9 open reading frame 114 (218565_at), score: 0.65 CA12carbonic anhydrase XII (203963_at), score: 0.46 CAND2cullin-associated and neddylation-dissociated 2 (putative) (213547_at), score: 0.44 CAPZA1capping protein (actin filament) muscle Z-line, alpha 1 (208374_s_at), score: 0.54 CBLBCas-Br-M (murine) ecotropic retroviral transforming sequence b (209682_at), score: -0.43 CCL11chemokine (C-C motif) ligand 11 (210133_at), score: -0.42 CCNT2cyclin T2 (204645_at), score: -0.47 CD320CD320 molecule (218529_at), score: 0.71 CD40CD40 molecule, TNF receptor superfamily member 5 (35150_at), score: -0.42 CD55CD55 molecule, decay accelerating factor for complement (Cromer blood group) (201925_s_at), score: -0.52 CDC42SE1CDC42 small effector 1 (218157_x_at), score: 0.51 CDCA4cell division cycle associated 4 (218399_s_at), score: -0.38 CDCP1CUB domain containing protein 1 (218451_at), score: -0.38 CDH18cadherin 18, type 2 (206280_at), score: 0.49 CDK5cyclin-dependent kinase 5 (204247_s_at), score: 0.45 CEP70centrosomal protein 70kDa (219036_at), score: 0.46 CHPcalcium binding protein P22 (214665_s_at), score: 0.43 CLCA2chloride channel accessory 2 (206165_s_at), score: -0.61 CLCF1cardiotrophin-like cytokine factor 1 (219500_at), score: -0.44 CLEC3BC-type lectin domain family 3, member B (205200_at), score: -0.63 CLEC4MC-type lectin domain family 4, member M (207995_s_at), score: -0.37 CLK1CDC-like kinase 1 (214683_s_at), score: -0.5 CLK4CDC-like kinase 4 (210346_s_at), score: -0.42 CLPBClpB caseinolytic peptidase B homolog (E. coli) (221845_s_at), score: 0.53 CLUclusterin (208791_at), score: -0.51 CLUAP1clusterin associated protein 1 (204576_s_at), score: 0.45 CNNM2cyclin M2 (206818_s_at), score: 0.45 CNPY3canopy 3 homolog (zebrafish) (217931_at), score: 0.43 COASYCoenzyme A synthase (201913_s_at), score: 0.42 COL14A1collagen, type XIV, alpha 1 (212865_s_at), score: -0.48 COL5A3collagen, type V, alpha 3 (52255_s_at), score: -0.69 COL7A1collagen, type VII, alpha 1 (217312_s_at), score: -0.46 COPS7BCOP9 constitutive photomorphogenic homolog subunit 7B (Arabidopsis) (219997_s_at), score: 0.45 COQ3coenzyme Q3 homolog, methyltransferase (S. cerevisiae) (221227_x_at), score: 0.5 CORO2Bcoronin, actin binding protein, 2B (209789_at), score: -0.59 CPT1Acarnitine palmitoyltransferase 1A (liver) (203633_at), score: 0.5 CR1complement component (3b/4b) receptor 1 (Knops blood group) (217484_at), score: 0.46 CRYL1crystallin, lambda 1 (220753_s_at), score: -0.41 CSADcysteine sulfinic acid decarboxylase (221139_s_at), score: -0.39 CST6cystatin E/M (206595_at), score: -0.54 CSTAcystatin A (stefin A) (204971_at), score: -0.38 CTSKcathepsin K (202450_s_at), score: -0.44 CTSOcathepsin O (203758_at), score: -0.5 CTSScathepsin S (202901_x_at), score: -0.38 CTSZcathepsin Z (210042_s_at), score: 0.55 CTTNcortactin (201059_at), score: 0.44 CXCL6chemokine (C-X-C motif) ligand 6 (granulocyte chemotactic protein 2) (206336_at), score: -0.46 CYLC2cylicin, basic protein of sperm head cytoskeleton 2 (207780_at), score: -0.38 CYP2A6cytochrome P450, family 2, subfamily A, polypeptide 6 (211295_x_at), score: -0.37 D4S234EDNA segment on chromosome 4 (unique) 234 expressed sequence (209569_x_at), score: -0.39 DARS2aspartyl-tRNA synthetase 2, mitochondrial (218365_s_at), score: 0.42 DCP2DCP2 decapping enzyme homolog (S. cerevisiae) (212919_at), score: -0.39 DCPSdecapping enzyme, scavenger (218774_at), score: 0.44 DIRAS3DIRAS family, GTP-binding RAS-like 3 (215506_s_at), score: -0.61 DPP4dipeptidyl-peptidase 4 (211478_s_at), score: -0.45 DUS4Ldihydrouridine synthase 4-like (S. cerevisiae) (205761_s_at), score: 0.53 DUSP5dual specificity phosphatase 5 (209457_at), score: -0.45 DUSP6dual specificity phosphatase 6 (208891_at), score: -0.5 ECM2extracellular matrix protein 2, female organ and adipocyte specific (206101_at), score: -0.38 EDNRBendothelin receptor type B (204271_s_at), score: -0.5 EGFL6EGF-like-domain, multiple 6 (219454_at), score: -0.38 EGLN3egl nine homolog 3 (C. elegans) (219232_s_at), score: -0.44 EIF2S1eukaryotic translation initiation factor 2, subunit 1 alpha, 35kDa (201143_s_at), score: -0.4 EIF4EBP2eukaryotic translation initiation factor 4E binding protein 2 (208770_s_at), score: -0.41 EMCNendomucin (219436_s_at), score: 0.59 EML1echinoderm microtubule associated protein like 1 (204797_s_at), score: -0.61 EML4echinoderm microtubule associated protein like 4 (220386_s_at), score: 0.45 EMP1epithelial membrane protein 1 (201325_s_at), score: -0.44 EPOerythropoietin (217254_s_at), score: 0.57 EREGepiregulin (205767_at), score: -0.45 ESM1endothelial cell-specific molecule 1 (208394_x_at), score: -0.4 ESRRAestrogen-related receptor alpha (1487_at), score: -0.45 EVI2Aecotropic viral integration site 2A (204774_at), score: 0.54 EVI2Becotropic viral integration site 2B (211742_s_at), score: 0.58 EXOSC5exosome component 5 (218481_at), score: 0.57 FAM134Bfamily with sequence similarity 134, member B (218532_s_at), score: -0.37 FAM174Bfamily with sequence similarity 174, member B (51158_at), score: -0.38 FAM63Bfamily with sequence similarity 63, member B (214691_x_at), score: -0.53 FAM70Afamily with sequence similarity 70, member A (219895_at), score: -0.41 FAM86Cfamily with sequence similarity 86, member C (220353_at), score: 0.63 FBXO31F-box protein 31 (219785_s_at), score: 0.56 FEM1Cfem-1 homolog c (C. elegans) (213341_at), score: -0.42 FGD2FYVE, RhoGEF and PH domain containing 2 (215602_at), score: -0.44 FKSG49FKSG49 (211454_x_at), score: 0.45 FLJ12529pre-mRNA cleavage factor I, 59 kDa subunit (217866_at), score: -0.4 FZD4frizzled homolog 4 (Drosophila) (218665_at), score: 0.6 GAAglucosidase, alpha; acid (202812_at), score: -0.51 GALTgalactose-1-phosphate uridylyltransferase (203179_at), score: 0.53 GANgigaxonin (220124_at), score: -0.56 GAS7growth arrest-specific 7 (202191_s_at), score: -0.38 GCAgrancalcin, EF-hand calcium binding protein (203765_at), score: -0.39 GKglycerol kinase (207387_s_at), score: -0.46 GNB1Lguanine nucleotide binding protein (G protein), beta polypeptide 1-like (220762_s_at), score: 0.66 GOLGA7golgi autoantigen, golgin subfamily a, 7 (217819_at), score: -0.38 GPM6Bglycoprotein M6B (209167_at), score: -0.49 GPR153G protein-coupled receptor 153 (221902_at), score: -0.48 GPR177G protein-coupled receptor 177 (221958_s_at), score: -0.46 GPR27G protein-coupled receptor 27 (221306_at), score: 0.44 GPRASP1G protein-coupled receptor associated sorting protein 1 (204793_at), score: 0.61 GPSM3G-protein signaling modulator 3 (AGS3-like, C. elegans) (204265_s_at), score: 0.56 GPSN2glycoprotein, synaptic 2 (208336_s_at), score: 0.48 GRLF1glucocorticoid receptor DNA binding factor 1 (202044_at), score: 0.42 GTF2H4general transcription factor IIH, polypeptide 4, 52kDa (203577_at), score: 0.48 HBBhemoglobin, beta (211696_x_at), score: -0.43 HERC3hect domain and RLD 3 (206183_s_at), score: -0.37 HGFhepatocyte growth factor (hepapoietin A; scatter factor) (209960_at), score: -0.59 HINFPhistone H4 transcription factor (206495_s_at), score: 0.45 HIST1H2AGhistone cluster 1, H2ag (207156_at), score: 0.43 HIST1H4Hhistone cluster 1, H4h (208180_s_at), score: -0.43 HSPB2heat shock 27kDa protein 2 (205824_at), score: 0.57 HTN1histatin 1 (206639_x_at), score: -0.38 HTR2A5-hydroxytryptamine (serotonin) receptor 2A (207135_at), score: -0.42 IER2immediate early response 2 (202081_at), score: -0.42 IGF2insulin-like growth factor 2 (somatomedin A) (202409_at), score: -0.52 IL13RA2interleukin 13 receptor, alpha 2 (206172_at), score: -0.5 IL33interleukin 33 (209821_at), score: -0.74 INTS8integrator complex subunit 8 (218905_at), score: 0.44 IPPKinositol 1,3,4,5,6-pentakisphosphate 2-kinase (219092_s_at), score: -0.38 IRF6interferon regulatory factor 6 (202597_at), score: -0.38 KAL1Kallmann syndrome 1 sequence (205206_at), score: -0.38 KCNE4potassium voltage-gated channel, Isk-related family, member 4 (222379_at), score: 0.51 KCNJ15potassium inwardly-rectifying channel, subfamily J, member 15 (210119_at), score: -0.47 KCTD14potassium channel tetramerisation domain containing 14 (58916_at), score: 0.52 KIAA0415KIAA0415 (209912_s_at), score: 0.45 KIAA0586KIAA0586 (205631_at), score: 0.56 KIAA1024KIAA1024 (215081_at), score: -0.42 KIRRELkin of IRRE like (Drosophila) (220825_s_at), score: 0.5 KLHL5kelch-like 5 (Drosophila) (220682_s_at), score: 0.43 KRT33Akeratin 33A (208483_x_at), score: -0.49 KTELC1KTEL (Lys-Tyr-Glu-Leu) containing 1 (218587_s_at), score: 0.43 LANCL2LanC lantibiotic synthetase component C-like 2 (bacterial) (218219_s_at), score: 0.47 LEMD3LEM domain containing 3 (218604_at), score: -0.38 LMAN2Llectin, mannose-binding 2-like (221274_s_at), score: 0.5 LOC100128223hypothetical protein LOC100128223 (221264_s_at), score: 0.55 LOC388152hypothetical LOC388152 (220602_s_at), score: 0.54 LOC390940similar to R28379_1 (213556_at), score: 0.43 LOC92249hypothetical LOC92249 (212957_s_at), score: 0.43 LRP1low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) (200785_s_at), score: -0.44 LRP2low density lipoprotein-related protein 2 (205710_at), score: -0.54 LY75lymphocyte antigen 75 (205668_at), score: -0.42 MAN1B1mannosidase, alpha, class 1B, member 1 (65884_at), score: -0.37 MAP1Smicrotubule-associated protein 1S (218522_s_at), score: -0.4 MAP4K4mitogen-activated protein kinase kinase kinase kinase 4 (218181_s_at), score: -0.38 MAP7D3MAP7 domain containing 3 (219576_at), score: 0.5 MASP2mannan-binding lectin serine peptidase 2 (207041_at), score: -0.41 MAST2microtubule associated serine/threonine kinase 2 (211593_s_at), score: -0.41 MBmyoglobin (204179_at), score: -0.39 MEGF6multiple EGF-like-domains 6 (213942_at), score: -0.39 MID1IP1MID1 interacting protein 1 (gastrulation specific G12 homolog (zebrafish)) (218251_at), score: -0.43 MLPHmelanophilin (218211_s_at), score: 0.58 MMP16matrix metallopeptidase 16 (membrane-inserted) (207012_at), score: -0.44 MSTP9macrophage stimulating, pseudogene 9 (213382_at), score: -0.41 MTHFD2Lmethylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2-like (220346_at), score: -0.46 MTHFSDmethenyltetrahydrofolate synthetase domain containing (218879_s_at), score: 0.5 MTMR1myotubularin related protein 1 (213511_s_at), score: -0.38 MTMR9myotubularin related protein 9 (204837_at), score: -0.46 MUC3Bmucin 3B, cell surface associated (214898_x_at), score: 0.5 MUSKmuscle, skeletal, receptor tyrosine kinase (207633_s_at), score: -0.46 MXRA5matrix-remodelling associated 5 (209596_at), score: -0.4 N6AMT1N-6 adenine-specific DNA methyltransferase 1 (putative) (220311_at), score: 0.43 NAGAN-acetylgalactosaminidase, alpha- (202943_s_at), score: 0.43 NAGPAN-acetylglucosamine-1-phosphodiester alpha-N-acetylglucosaminidase (205090_s_at), score: 0.52 NAT13N-acetyltransferase 13 (GCN5-related) (217745_s_at), score: -0.41 NDUFA2NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 2, 8kDa (213550_s_at), score: 0.52 NEFLneurofilament, light polypeptide (221805_at), score: -0.61 NELFnasal embryonic LHRH factor (221214_s_at), score: -0.4 NFATC2IPnuclear factor of activated T-cells, cytoplasmic, calcineurin-dependent 2 interacting protein (217526_at), score: -0.61 NMBneuromedin B (205204_at), score: 0.59 NME6non-metastatic cells 6, protein expressed in (nucleoside-diphosphate kinase) (205851_at), score: 0.43 NOL8nucleolar protein 8 (218244_at), score: 0.5 NOS1nitric oxide synthase 1 (neuronal) (207309_at), score: 0.45 NR0B1nuclear receptor subfamily 0, group B, member 1 (206644_at), score: -0.39 NR5A2nuclear receptor subfamily 5, group A, member 2 (208343_s_at), score: -0.41 NSUN5BNOL1/NOP2/Sun domain family, member 5B (214100_x_at), score: 0.43 NTHL1nth endonuclease III-like 1 (E. coli) (209731_at), score: 0.49 NTNG1netrin G1 (206713_at), score: -0.47 NUDT1nudix (nucleoside diphosphate linked moiety X)-type motif 1 (204766_s_at), score: 0.49 NUDT6nudix (nucleoside diphosphate linked moiety X)-type motif 6 (220183_s_at), score: 0.62 OAS12',5'-oligoadenylate synthetase 1, 40/46kDa (205552_s_at), score: -0.4 OLFM1olfactomedin 1 (213131_at), score: -0.46 OLFML2Aolfactomedin-like 2A (213075_at), score: -0.53 OPCMLopioid binding protein/cell adhesion molecule-like (214111_at), score: -0.41 PAFAH1B3platelet-activating factor acetylhydrolase, isoform Ib, gamma subunit 29kDa (203228_at), score: 0.53 PDE8Bphosphodiesterase 8B (213228_at), score: -0.39 PEG3paternally expressed 3 (209242_at), score: -0.41 PENKproenkephalin (213791_at), score: -0.44 PFDN6prefoldin subunit 6 (222029_x_at), score: 0.45 PHC3polyhomeotic homolog 3 (Drosophila) (215521_at), score: -0.43 PID1phosphotyrosine interaction domain containing 1 (219093_at), score: -0.41 PIGLphosphatidylinositol glycan anchor biosynthesis, class L (213889_at), score: 0.52 PKIAprotein kinase (cAMP-dependent, catalytic) inhibitor alpha (204612_at), score: -0.43 PLAURplasminogen activator, urokinase receptor (211924_s_at), score: -0.4 PLEKHA2pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 2 (217677_at), score: 0.47 PLXNB1plexin B1 (215807_s_at), score: -0.43 PMAIP1phorbol-12-myristate-13-acetate-induced protein 1 (204285_s_at), score: -0.53 PMS2CLPMS2 C-terminal like pseudogene (221206_at), score: -0.4 PNRC2proline-rich nuclear receptor coactivator 2 (217779_s_at), score: -0.49 POP1processing of precursor 1, ribonuclease P/MRP subunit (S. cerevisiae) (213449_at), score: 0.52 POU4F1POU class 4 homeobox 1 (206940_s_at), score: -0.55 PPFIBP1PTPRF interacting protein, binding protein 1 (liprin beta 1) (214374_s_at), score: 0.47 PRELPproline/arginine-rich end leucine-rich repeat protein (204223_at), score: -0.39 PRKG2protein kinase, cGMP-dependent, type II (207505_at), score: -0.72 PRSS2protease, serine, 2 (trypsin 2) (205402_x_at), score: -0.38 PRSS3protease, serine, 3 (213421_x_at), score: -0.51 PSD4pleckstrin and Sec7 domain containing 4 (203317_at), score: -0.4 PSMB4proteasome (prosome, macropain) subunit, beta type, 4 (202243_s_at), score: 0.5 PSMB9proteasome (prosome, macropain) subunit, beta type, 9 (large multifunctional peptidase 2) (204279_at), score: 0.44 PTCD1pentatricopeptide repeat domain 1 (218956_s_at), score: 0.54 PTENP1phosphatase and tensin homolog pseudogene 1 (217492_s_at), score: 0.45 PTGS1prostaglandin-endoperoxide synthase 1 (prostaglandin G/H synthase and cyclooxygenase) (215813_s_at), score: -0.55 PTGS2prostaglandin-endoperoxide synthase 2 (prostaglandin G/H synthase and cyclooxygenase) (204748_at), score: -0.4 PTMAprothymosin, alpha (200772_x_at), score: 0.44 PTPRBprotein tyrosine phosphatase, receptor type, B (205846_at), score: -0.4 PTTG3pituitary tumor-transforming 3 (208511_at), score: 0.47 RAB8BRAB8B, member RAS oncogene family (219210_s_at), score: 0.53 RALGDSral guanine nucleotide dissociation stimulator (209050_s_at), score: 0.48 RAPGEF6Rap guanine nucleotide exchange factor (GEF) 6 (219112_at), score: -0.39 RARRES1retinoic acid receptor responder (tazarotene induced) 1 (221872_at), score: -0.45 RASA2RAS p21 protein activator 2 (206636_at), score: -0.41 RASSF2Ras association (RalGDS/AF-6) domain family member 2 (203185_at), score: -0.48 RBM39RNA binding motif protein 39 (207941_s_at), score: -0.51 RCC1regulator of chromosome condensation 1 (206499_s_at), score: 0.47 RFX7regulatory factor X, 7 (218430_s_at), score: 0.57 RHBDD3rhomboid domain containing 3 (204402_at), score: -0.48 RNF2ring finger protein 2 (205215_at), score: -0.45 RNF44ring finger protein 44 (203286_at), score: 0.52 RP2retinitis pigmentosa 2 (X-linked recessive) (205191_at), score: -0.48 RPL14ribosomal protein L14 (219138_at), score: 0.57 RPL23ribosomal protein L23 (214744_s_at), score: -0.4 RPL27Aribosomal protein L27a (212044_s_at), score: 0.65 RPL38ribosomal protein L38 (202028_s_at), score: 0.48 RPLP2ribosomal protein, large, P2 (200908_s_at), score: 0.43 RPS11ribosomal protein S11 (213350_at), score: 0.66 SAT1spermidine/spermine N1-acetyltransferase 1 (213988_s_at), score: -0.57 SEC14L5SEC14-like 5 (S. cerevisiae) (210169_at), score: -0.47 SERPINB7serpin peptidase inhibitor, clade B (ovalbumin), member 7 (206421_s_at), score: 0.45 SERPINB9serpin peptidase inhibitor, clade B (ovalbumin), member 9 (209723_at), score: 0.55 SETMARSET domain and mariner transposase fusion gene (206554_x_at), score: 0.68 SEZ6L2seizure related 6 homolog (mouse)-like 2 (218720_x_at), score: -0.46 SFRS12splicing factor, arginine/serine-rich 12 (212721_at), score: -0.46 SIK2salt-inducible kinase 2 (213221_s_at), score: -0.41 SLC16A5solute carrier family 16, member 5 (monocarboxylic acid transporter 6) (206600_s_at), score: 0.55 SLC16A6solute carrier family 16, member 6 (monocarboxylic acid transporter 7) (207038_at), score: -0.39 SLC17A7solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 7 (204230_s_at), score: -0.47 SLC20A2solute carrier family 20 (phosphate transporter), member 2 (202744_at), score: -0.42 SLC25A17solute carrier family 25 (mitochondrial carrier; peroxisomal membrane protein, 34kDa), member 17 (211754_s_at), score: -0.43 SLC38A4solute carrier family 38, member 4 (220786_s_at), score: 0.61 SLKSTE20-like kinase (yeast) (206875_s_at), score: -0.47 SMU1smu-1 suppressor of mec-8 and unc-52 homolog (C. elegans) (218393_s_at), score: 0.51 SNHG3small nucleolar RNA host gene 3 (non-protein coding) (215011_at), score: 0.42 SNORA21small nucleolar RNA, H/ACA box 21 (215224_at), score: 0.42 SNX13sorting nexin 13 (213292_s_at), score: 0.52 SORT1sortilin 1 (212807_s_at), score: 0.49 SOX9SRY (sex determining region Y)-box 9 (202935_s_at), score: -0.43 SPA17sperm autoantigenic protein 17 (205406_s_at), score: 0.54 SPAM1sperm adhesion molecule 1 (PH-20 hyaluronidase, zona pellucida binding) (216989_at), score: -0.38 SPP1secreted phosphoprotein 1 (209875_s_at), score: 0.55 STC1stanniocalcin 1 (204597_x_at), score: -0.41 STK16serine/threonine kinase 16 (209622_at), score: 0.48 STOMstomatin (201060_x_at), score: 0.43 SYNMsynemin, intermediate filament protein (212730_at), score: 0.51 SYT1synaptotagmin I (203999_at), score: -0.47 SYT11synaptotagmin XI (209198_s_at), score: 0.55 TAF9BTAF9B RNA polymerase II, TATA box binding protein (TBP)-associated factor, 31kDa (221618_s_at), score: 0.44 TBX5T-box 5 (207155_at), score: 0.45 TCF7L1transcription factor 7-like 1 (T-cell specific, HMG-box) (221016_s_at), score: 0.42 TFDP2transcription factor Dp-2 (E2F dimerization partner 2) (203588_s_at), score: 0.44 TGFAtransforming growth factor, alpha (205016_at), score: -0.38 TGFBRAP1transforming growth factor, beta receptor associated protein 1 (205210_at), score: 0.54 THOC2THO complex 2 (212994_at), score: 0.5 THOC6THO complex 6 homolog (Drosophila) (218848_at), score: 0.47 THYN1thymocyte nuclear protein 1 (218491_s_at), score: 0.46 TIMM9translocase of inner mitochondrial membrane 9 homolog (yeast) (218316_at), score: 0.49 TMEM158transmembrane protein 158 (213338_at), score: -0.42 TMEM160transmembrane protein 160 (219219_at), score: 0.55 TMSB15Athymosin beta 15a (205347_s_at), score: 0.65 TOMM22translocase of outer mitochondrial membrane 22 homolog (yeast) (217960_s_at), score: 0.52 TOXthymocyte selection-associated high mobility group box (204529_s_at), score: 0.45 TP53TG1TP53 target 1 (non-protein coding) (209917_s_at), score: 0.5 TPK1thiamin pyrophosphokinase 1 (221218_s_at), score: -0.66 TRAPPC10trafficking protein particle complex 10 (209412_at), score: -0.38 TRIAP1TP53 regulated inhibitor of apoptosis 1 (218403_at), score: 0.45 TRIB2tribbles homolog 2 (Drosophila) (202478_at), score: 0.43 TRMT61BtRNA methyltransferase 61 homolog B (S. cerevisiae) (221229_s_at), score: 0.44 TRPC1transient receptor potential cation channel, subfamily C, member 1 (205802_at), score: -0.49 TRPV4transient receptor potential cation channel, subfamily V, member 4 (219516_at), score: -0.37 UCHL5IPUCHL5 interacting protein (213334_x_at), score: 0.44 UCP2uncoupling protein 2 (mitochondrial, proton carrier) (208998_at), score: 1 UFM1ubiquitin-fold modifier 1 (218050_at), score: -0.42 USE1unconventional SNARE in the ER 1 homolog (S. cerevisiae) (221706_s_at), score: 0.46 USP47ubiquitin specific peptidase 47 (221518_s_at), score: 0.45 WDR43WD repeat domain 43 (214662_at), score: 0.44 WDR52WD repeat domain 52 (221103_s_at), score: -0.41 WRNWerner syndrome (205667_at), score: -0.4 XPO4exportin 4 (218479_s_at), score: 0.46 YIPF1Yip1 domain family, member 1 (214733_s_at), score: 0.43 YY2YY2 transcription factor (216531_at), score: 0.55 ZMAT5zinc finger, matrin type 5 (218752_at), score: 0.49 ZNF124zinc finger protein 124 (206928_at), score: 0.43 ZNF174zinc finger protein 174 (210291_s_at), score: 0.52 ZNF195zinc finger protein 195 (204234_s_at), score: 0.46 ZNF219zinc finger protein 219 (219314_s_at), score: -0.56 ZNF22zinc finger protein 22 (KOX 15) (218005_at), score: 0.43 ZNF236zinc finger protein 236 (47571_at), score: -0.43 ZNF324Bzinc finger protein 324B (222318_at), score: -0.4 ZNF350zinc finger protein 350 (219266_at), score: 0.49 ZNF667zinc finger protein 667 (207120_at), score: 0.56 ZNF749zinc finger protein 749 (215289_at), score: -0.46 ZRANB1zinc finger, RAN-binding domain containing 1 (201219_at), score: 0.44
| Id | sample | Experiment | ExpName | Array | Syndrome | Cell.line |
|---|---|---|---|---|---|---|
| E-TABM-263-raw-cel-1515486011.cel | 19 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486391.cel | 38 | 6 | Cycle | hgu133a2 | none | Cycle 1 |
| E-TABM-263-raw-cel-1515486191.cel | 28 | 6 | Cycle | hgu133a2 | none | Cycle 1 |