In this paper, Molly K. Burke and his collogues did an experimental evolution systems, which allows the genomic study of adaptation. They selected outbred, sexually reproducing, replicated populations of Drosophila melanogaster, which experienced over 600 generations of laboratory selection for accelerated development.
Short-read sequences from three genomic DNA libraries, were obtained using Illumina platform, they are as follows:
a) A pooled sample of five replicate populations that have undergone sustained selection for accelerated development and early fertility for over 600 generations (ACO);
b) A pooled sample of five replicate ancestral control populations, which experience no direct selection on development time (CO);
c) A single ACO replicate population (ACO1);
 |
| Figure 1: Summary of phenotypic divergence in the selection treatments. |
In the above figure, the grey bar indicates values measured in the ACO and CO treatments for each of the five replicate populations. B indicates replicate populations, which represent phenotypes typical of populations kept on two-week generation maintenance schedules.
This figure shows a comparative analysis between the ACO population and the population with the CO treatment. Every time, ACO featured significantly differentiated phenotypes, including shorter development time and reductions in pre-adult viability, longevity, adult body size and stress resistance. Furthermore, the CO treatment does not show stringent selection, as it entails no more than moderate selection for postponed reproduction, resulting in moderately increased development time and longevity.
 |
|
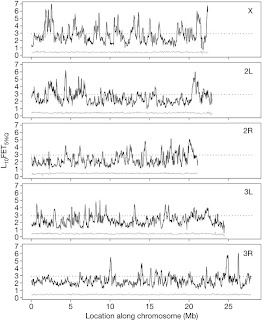
Figure 2: Differentiation throughout the genome.
|
A 100-kb genome-wide sliding-window analysis test was carried out to identify regions diverged in allele frequency between the ACO and CO libraries and between the ACO and ACO1 libraries. This is due to the fact that, linkage disequilibrium in individual ACO and CO replicate populations may extend anywhere from 20 to 100 (kb). The five different panels are basically for the five major D. melanogaster chromosome arms.
It identifies a large number of genomic regions showing significant divergence between the accelerated development populations and their matched controls depicted by black lines in Figure 2, and very little divergence is observed between a single replicate evolved population (ACO1) and the pooled sample consisting of all five ACO populations and this is indicated by the grey lines. The dotted line is the threshold that any given window has a 0.1% chance of exceeding relative to the genome-wide level of noise. Interestingly, an excess of diverged regions on the X chromosome relative to on the autosomes is very evident. This observation is only expected if adaptation were driven by selection on initially rare recessive or partially recessive alleles. Furthermore, the sharpness of the peaks suggests that regions of the genome that have responded to experimental evolution are precisely identified. However, even the sharpest peaks tend to delineate, 50–100-kb regions.
Kaplan et al. stated that, recent research on evolutionary genetics has focused on classic selective sweeps. In a recombining region, a selected sweep is expected to reduce heterozygosity at SNPs flanking the selected site.
 |
| Figure 3: Heterozygosity throughout the genome |
Figure 3, shows a similar sliding-window analysis (100 kb) of heterozygosity in ACO and CO lines suggesting that there are indeed local losses of heterozygosity, which is depicted by the red and the blue lines, respectively. Heterozygosity in ACO1 depicted by grey line, shows remarkable concordance with the reductions in heterozygosity in the ACO pool. Regions of reduced heterozygosity are strongly associated with regions of differentiated allele frequency. Interestingly, we observed no location in the genome where heterozygosity is reduced to anywhere near zero, and therefore, it lacks the evidence for a classic sweep is a feature of the data regardless of window size. Nevertheless, both the figure 2 and 3 are quite similar.
 |
|
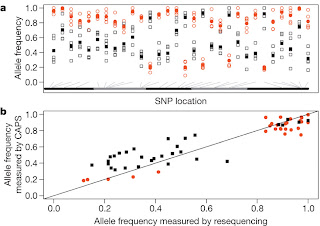
Figure 4: Analysis of individual genotypes, measured by cleaved amplified
polymorphic sequence (CAPS) techniques.
|
In Figure 4a, allele frequency estimates of the most common allele at 30 SNPs genotyped in 35 females per replicate population. Red circles and grey squares represent ACO and CO estimates. Open symbols are allele frequencies for ACO1–ACO5 and CO1–CO5, and filled symbols represent treatment means. Alternating black and grey bars designate the X, 2L, 2R, 3L, and 3R arms, respectively. The grey lines indicate SNP location. We observe that replicate populations within a selection treatment have very similar allele frequencies.
In Figure 4b, it gives a scatter plot comparing allele frequency estimates at the same 30 SNPs obtained from the Illumina resequencing versus individual genotyping. Red circles represent ACO, black squares represent CO and the straight line represents a slope of unity. Here we see that individual genotypes are consistent with allele frequency estimates from the resequenced pooled libraries.
Therefore, it can be concluded that the congruence in allele frequencies and patterns of heterozygosity between the ACO1 and ACO libraries is unlikely to be some sort of artefact of sample preparation or data analysis.
The study shows a convergence of allele frequencies and heterozygosity levels between replicate populations. This convergence might be due to selection, acting on the same intermediate-frequency variants in each population. Under this scenario, convergence in allele frequencies is due to parallel evolution. Another reason could be, unwanted migration between replicate populations, even at very low levels.
Conclusively, it was very interesting to see that despite strong selection, Molly K. Burke and his collogues failure to observe the signature of a classic sweep in these populations.
